Identification of Resistance Sources Among Traditional Cotton Genotypes from Burkina Faso Against Xanthomonas Citri Pv. Malvacearum
Wonni Issa1*, Bourgou Larbouga1, Tarpaga Vianney1, Sanga Issouf2 and Ouedraogo Leonard1
1centre National de Recherche Scientifique et Technologique (CNRST)/Institut de l’Environnement et de Recherches Agricoles (INERA), Burkina Faso, Africa
2Université Nazi BONI, Burkina Faso, Africa
Submission: August 12, 2019; Published: October 28, 2019
*Corresponding author: Wonni Issa, Centre National de Recherche Scientifique et Technologique (CNRST)/Institut de l’Environnement et de Recherches Agricoles (INERA), Burkina Faso, Africa
How to cite this article: Wonni Issa, Bourgou Larbouga, Tarpaga Vianney, Sanga Issouf, Ouedraogo Leonard. Identification of Resistance Sources Among Traditional Cotton Genotypes from Burkina Faso Against Xanthomonas Citri Pv. Malvacearum. Agri Res& Tech: Open Access J. 2019; 23(2): 556226. DOI: 10.19260/ARTOAJ.2019.22.556226
Abstract
Bacterial blight of Cotton caused by Xanthomonas citri. pv malvacearum, is a severe disease occurring in all cotton-growing areas. In several countries, control has been achieved through the use of resistant cotton genotypes associated with agricultural methods limiting disease spread. The main objective of the present study was to identify resistance sources against races 18 and 20 of Xanthomonas citri pv. malvacearum in artificial inoculation conditions within a collection of local cotton cultivars. Our results show that two conventional and commercialized varieties, FK64 and FK37, and traditional accessions belonging to the species hirsutism, barbadense and arborerum harbor resistance against X. citri pv. malvacearum race 18 from leaves, stem and boll inoculation. However, with race 18 strain, some cultivars conserve bacteria into stem and leave that remain asymptomatic 35 days after inoculation. No source of resistance was detected against race 20 of X. citri pv. malvacearum. For better management of cotton blight, the knowledge of the phenotypic diversity of pathogen strains, their spatial and temporal distribution is a prerequisite to guide the use of resistance gene in breeding program and their deployment in cotton growing areas in Burkina Faso.
Keywords: Gossypium ssp, Xanthomonas citri. pv malvacearum, Cotton Bacterial Blight, Burkina Faso
Introduction
Gossypium sp., belonging to Malvaceae family, represents is the most widely grown fiber plant in the world. It is native to tropical and subtropical regions around the world, including America, India and Africa. However, almost all the commercial cotton grown in the world comes from varieties of the native species Gossypium hirsutum and G. barbadense. In Africa, Mali is the largest producer, followed by Burkina Faso with a production of 613,000 tons for the season 2017-2018. Despite its strong contribution to youth employment and rural people’s incomes, the cotton farming faces many parasitic constraints including cotton bacterial blight (CBB) disease caused by Xanthomonas citri pv. malvacearum (Xcm). The first detailed description of CBB was made in the US [1]. Today, this disease occurs in all cotton-growing areas throughout the world [2,3] and was shown to be a serious limiting factor of fiber production in the US [4], India [5] and in Africa [6]. Xanthomonas citri pv. malvacearum infects plant tissues and organs of cotton during all stages of development beginning with the seedling stage [1]. Foliar disease is characterized by angular leaf spot with a red to brown border. Leaf petioles and stems may also become infected (cotton black leg), which can lead to premature defoliation of the plant. Infected bolls have round, as opposed to angular, lesions that initially may appear water-soaked, and then become sunken and dark brown to black. Bolls infected with Xanthamonas may result in rotted seed and discolored lint. Under bacterial blight infection, boll yield losses up to 35% have been reported in India. In Pakistan, losses up to 50% of yield have been estimated by [7]. However, in West Africa and particularly in Burkina Faso, data on production losses are not available. Given the favorable climate for development CBB (hot and humid) and the apparition of new races of Xcm, it can be assumed that equivalent levels of losses to those observed in other regions of the world are recorded in Africa [8]. Indeed, the seed is the main source of inoculum necessary for the start of the infectious process of the aerial organs. According several authors, most lines resistant to Xcm exist in G. hirsutum cultivars [9,10,11,12]. To date, 22 races have been reported [13]. However, at least 18 genes has been identified and participate in the breeding for the resistance against Xcm [2,13]. Most races are geographically distinct. In the U.S, race 18 is predominant [12]. InBurkina Faso, Chad and Sudan, [8] reported that in 1981, news strains of Xcm (race 20) overcame the resistance of varieties with the B2-B3 and B91-B101 genes that express a total leaf resistance to CBB. Since then, resistance to bacterial blight is an important parameter in breeding programs. The objective of our study is to screen the local cotton accessions from Burkina Faso in order to identify sources of resistance against the races 18 and 20 of Xcm.
Materials and Methods
Plant Material

The plant material of this study is part of the 358 accessions collected in different agro-ecological areas of Burkina Faso. Previous characterizations indicated that all the accessions are shared between four different species, two tetraploids G. hirsutum and G. barbadense and two diploids G. arboreum and G. herbaceum [14,15]. A subset of 15 accessions (6 G. arboreum, 5 G. barbadense and 4 G. hirsutum) was used for our current study. Three popularized G. hirsutum cultivars (FK37, FK64 and Stam 59A), were also included as controls. A detailed list of all the genotypes investigated is presented in Table 1. The origin of local cultivars is the producer who given the seed and the region where it was collected.
Xanthomonas citri pv. Malvacearum strains
Two X. citri pv. malvacearum strains representing race 18 (X18) and race 20 (X20) originated from Burkina Faso, were used to screen the different cultivars. The race 20 was identified as the high virulent strain in cotton growing areas in Africa [16].
Inoculation Methods
The method used to inoculate the different organs of the cotton plant was related to the development stage and the type of organ (Leaf, stem, flower and boll). All the infections were performed using bacterial suspension concentrated to 108 CFU/ ml. Leaf infection was performed using the modified Sandpaper method described by [17]. The leaves were placed between cotton soaked in the bacterial suspension containing carborandum and light pressure was applied. About the stem, the bacterial suspension was infiltrated under the five terminal leaves using a needle syringe. Then, the inoculation site was plugged with parafilm to prevent external infection. On the flower buds and capsules, the inoculation was performed using the needle syringe by direct infiltration on the floral peduncle and into the capsule. After inoculation, the plants were watered and covered with a polythene bag for 48 hours.
Evaluation of Cotton Genotypes for CBB Resistance
Disease incidence was assessed by the presence/absence of typical leaf symptoms at 7 days after inoculation as follow:

The severity of foliar disease (DS) was recorded weekly by using the scale of [18] (Table 2). SD values were used to calculate AUDPC using the formulas described by [19] and [20]as follows:
SD (%) = Σ (number of diseased plants in a rating category × that rating category) / (total number of plants × category maximum score) × 100

where xi is CBB severity at the time i , n is the number of data taken and t is the number of days between the registration of xi and x1=i .
On flower buds and bolls, the incidence (%) was evaluated at 15 DAI by counting respectively the number of flower buds and aborted bolls in relation to the total number of each organ inoculated. In addition, the diameter of the canker or rot induced by inoculation was measured.

Xcm Survival in Cotton Plant
To verify the survival capacity of the bacteria into cotton plant at 30 JAI, Xcm were re-isolated on culture media respectively from stems and leaves. The presence (+) or absence (-) of bacteria into tissues was recorded.
Data Analysis
All data were recorded using Excel 2013. An ANOVA was performed with the SISVAR 5.1 Build 72 software. The means of incidence and AUDPC were compared using the Scott-Knott test, based on a 5% threshold.
Results
Disease Incidence and Symptoms
After inoculation, various symptoms were observed on the different part of plant (Figure 1), but with varying incidence levels according to cultivars and Xcm races. Thus, on the leaves, we found symptoms of bacterial wilt, hypersensitivity reactions and even resistance reactions especially with race 18. On the stems, all inoculated genotype showed cankers or rot but at different lengths (Figure 2). Cultivars Ec214, PE28, Ec51, Ec322, Ec48 and Ec235 were less susceptible to stem canker with length less to 2 cm. On the flower buds, we observed 100% abortion with race 20 independently to genotype (Table 3). Some flowers that did not aborted evolved to give boll. However, these boll, in comparison with control were not good quality. We observed dropping of bolls, but also rots of boll with different diameters according to genotype and the both Xcm strain (Table 3). Our results show that all cultivars were susceptible to the both races at flower and boll inoculation.


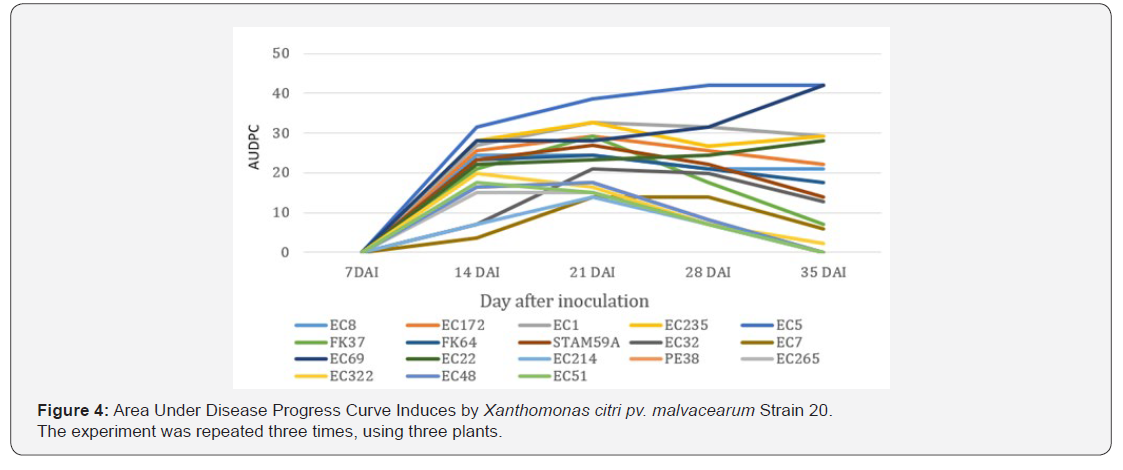
CBB Severity
Figures 3 & 4 show the weekly evolution of the disease caused by races 18 and 20 respectively. AUDPCs calculated on a five-week post-inoculation observation showed significant differences (P ≤ 0.05) between cotton genotypes and races. The first symptoms appear 7 days after inoculation of both races. With race 18, CBB evolves for all varieties up to three weeks after inoculation, excepted FK37 and EC22 genotypes that showed immunity against leaf infection. However, with the genotypes FK64, STAM59A, EC7, EC214, PE38, EC265, EC322, EC48 and EC51, the severity of CBB decreases 3 weeks after inoculation and disappears after 5 weeks after inoculation. With race 20, all genotypes were susceptible with a maximum level of severity recorded three weeks after inoculation. At 35 DAI, only five genotypes EC214, PE38, EC265, EC48, EC51 showed symptoms of CBB.
Survival Ability of Xcm into Cotton Tissues
The results showed that the bacteria are more conserved in the stem than in the leaves whatever the variety (Data not showed). In general, 30 DAI, bacteria are absent into last five terminal leaves excepted cultivars PE38, EC69 and Ec7 that X18 was detected. In the stems, the inoculation site remains infest until 30 JAI. We observed that the bacteria progress more in the part above the point of inoculation. Race 20 caused stem rot on some cultivars leading to plant wilting.
Discussion

Evaluation of 18 cotton cultivars from Burkina Faso, showed that some of them hold the full potential to control race18 but not race 20 of Xanthomonas citri pv. malvacearum, the causal agent of cotton bacterial blight. However, the level of susceptibility varies according to the plant parts that are the leaves, stems, flowers and capsules. According [20,21,22] resistance to CBB varied greatly within the genus Gossypium. Cultivars FK37 and EC22, inoculated with race 18, the resistance response induced at the site of inoculation, look like an HR-like phenotype. Then, Ec32 and FK64 are resistant against X18. Interestingly, these cultivars, immune or resistant against race 18, have the lowest flower buds abortion rates. In addition, cultivars FK64 and FK37 are moderately resistant to boll infection. These results show a high resistance of some cultivars from G. barbadense against CBB although little resistance occurred naturally in this specie [23]. Our results confirm that genotypes from G. hirsutum provided the broadest spectrum of disease expression, varying from fully susceptible to resistant. Indeed, Ec22 and Ec32 are resistant while STAM59A and Ec69 are highly susceptible to race 18. Indeed, [24], reported that no single R gene confers durable resistance or immunity to CBB, due to evolutionary shifts in Xcm virulence in response to the selection pressures imposed by resistant varieties. However, pyramiding of B genes has been sometimes successfully used to produce lines with high resistance against multiple races of Xcm. For example, pyramiding B2 with B3 and other polygenic complexes [25], has provided substantial protection against all races of Xcm identified in the US [26]. In Africa, resistance to CBB was obtained by combining B genes (B2B3 and B9LB10 L) from G. hirsutum which confer resistance to all Xcm races except races 20, 21 and 22 [13]. In a concordant way, our study reveal that, no cultivar confer a resistance to race 20. This race attacks all cultivars with an important symptoms on the leaves, stems, flowers, bolls whatever the species of cotton. Other interesting results is that bacteria survive more time in the stem than the leaves after inoculation. In contrast, bacteria were detected from un-inoculated leaves without symptoms following the stem infection. In fact, these leaves would be infected following the production of secondary inoculum either from the stem or from leaf exudates. In addition to these sources, previous studies indicate that asymptomatic winter weeds remaining in field could potentially serve as a source of CBB inoculum the following season [27]. Therefore, any management practice to promote drying of cotton leaves and stems may help reduce the development of CBB. Except STAM56A, the both other conventional varieties marketed that are FK37 and FK64, resist to race 18. In recent years, although the incidence of CBB is great in some cotton growing areas, the economic damage are weak and suggest that the strains or races of Xcm are less virulent. However, the introduction or broad propagation of race 20 already identified in Burkina Faso or/and another virulent race would be damage for cotton production. As a preventive measure, it is essential to map the Xcm races and follow their dynamics in order to anticipate any new CBB epidemic.

Conclusion
Our study confirms the presence of bacterial cotton disease in Burkina Faso. The highest prevalence of the disease has been observed in the Faso Cotton and SOCOMA zones. Among cultivars and conventional varieties, cultivar EC-14 was resistant to collar infection with races 18 and 20; while the cultivar EC22 and the conventional FK37 variety were resistant to leaf disease inoculated with race 18. However, against race 20, G. arboreum genotypes were found to be more susceptible than conventional varieties.
Acknowledgments
This research was supported by the three cotton societies of Burkina Faso which SOFITEX, Faso Coton and SOCOMA.
References
- Atkinson GF (1891) The black rust of cotton. Coop. Ext. Serv, Alabama Agricultural Experiment Station Bulletin 27, Univ. of Alabama, Tuscaloosa, USA.
- Hillocks R (1992) Bacterial blight. In: Cotton diseases. Wallingford, CAB International p. 39-85.
- Zomorodian A, Rudolph K (1993) Xanthomonas campestris pv. malvacearum, cause of bacterial blight of cotton. In: Xanthomonas by Swings JG, Civerolo EL, Chapman (Eds.), Hall London, UK, p. 25-30.
- Senchina DS, Alvarez I, Cronn RC, Liu B, Rong J et al. (2003) Rate variation among nuclear genes and the age of polyploidy in Gossypium. Molecular Biology and Evolution 20: 633-643.
- Verma JP (1986) Bacterial Blight of Cotton. Boca Raton. CRC Press Inc FL, USA.
- Follin JC, Girardot B, Mangano V, Benetez R (1988) Nouveaux résultats sur le déterminisme génétique de la résistance foliaire totale du cotonnier (Gossypium hirsutum) à la bactériose Xanthomonas campestris pv. malvacearum (Smith) Dye, races 18 et 20. Cot. et Fib. Trop 43: 167-175.
- Huissain T, Ali M (1975) A review of cotton diseases of Pakistan. Pakistan cotton 19(2): 71-86.
- Ouédraogo SL, Sanfo D, Somda I, Tiemtore BC (2009) Analyse de l’influence du fonds génétique, des conditions climatiques et du mode de protection phytosanitaire sur l’expression de la bactériose chez différentes variétés de cotonnier au Burkina Faso. Tropicultura 27:31-34.
- Knight R, Clouston T (1939) The genetics of blackarm resistance. J Genet 38(1): 133-59.
- Bourland F, Myers GO (2015) Conventional cotton breeding. In: Cotton. American Society of Agronomy, Inc., Crop Science Society of America, Inc., and Soil Science Society of America, Inc pp. 205-228.
- Bourland FM, Jones DC (2015) Registration of Arkot 0305. Arkot 0306. Arkot 0309. and Arkot 0316 germplasm lines of cotton. J Plant Registrations 9(1): 94-98.
- Showmaker KC, Mark AA, Chuan YH, Brigitte EM, Xiaoqiang W, et al. (2017) The genome of the cotton bacterial blight pathogen Xanthomonas citri pv. malvacearum strain MSCT1. Standards in Genomic Sciences 12: 42.
- Delannoy E, Lyon BR, Marmey P, Jalloul A, Daniel JF, et al. (2005) Resistance of cotton towards Xanthomonas campestris pv. malvacearum. Annu Rev Phytopathol 43: 63-82.
- Bourgou L, Tarpaga WV, Sanfo D, Sawadogo M, Zongo JD (2014) Préservation de la diversité génétique dans le genre Gossypium au Burkina Faso: collecte et évaluations. Int. J. Biol. Chem Sci 8(5): 2081-2094.
- Bourgou L, Sawadogo M, Sanfo D, Lacape JM (2016) SSR-based genetic diversity of traditional and perennial cotton (Gossypium spp.) populations collected in Burkina Faso. Genet Resour Crop Evol 64(7): 1743-1759.
- Huang X, Zhai J, Luo Y, Rudolph K (2008). Identification of a highly virulent strain of Xanthomonas axonopodis pv. malvacearum. European Journal of Plant Pathology 122(4): 461-469.
- Eddin KS, Marimuthu T, Ladhalakshmi D, Rabindran R, Velazhahan R (2005) A simple inoculation technique for evaluation of cotton genotypes for resistance to bacterial blight caused by Xanthomonas axonopodis pv. malvacearum. Journal of Plant Diseases and Protection 112 (4): 321–328.
- Yehouessi MT (1988) Protocole de cotation bactériose campagne 1987/1988. S.R.C.F.J. Station de N’Taria B.P. 28. Koutiala. Cellule génétique p. 3.
- Bansal VK, Kharbanda PD, Stringam GR, Thiagarajah MR, Tewari JP (1994) A Comparison of Greenhouse and Field Screening Methods for Blackleg Resistance in Doubled Haploid Lines of Brassica-Napus. Plant Disease 78: 276-281.
- Mamba MG, tshilenge DP, Nkongolo KK, Kalonji MA (2014) Characterization of Congolese Strains of Xanthomonas axonopodis pv. manihotis Associated with Cassava Bacterial Blight. American Journal of Plant Sciences 5: 1191-1201.
- Hunter RE, Brinkerhoff LA, Bird LS (1968) Development of a set of Upland cotton lines for differentiating races of Xanthomonas malvacearum. Phytopathology 58: 830-832.
- Wallace TP, El-Zik KM (1989) Inheritance of resistance in three cotton cultivars to the HV1 isolate of bacterial blight. Crop Science pp. 1114-1119.
- Jalloul A, Sayegh M, Champion A, Nicole M (2015) Bacterial blight of cotton. Phytopathologia Mediterranea 54(1): 3-20.
- Brinkerhoff LA, Verhalen LM, Johnson WM, Essenberg M, Richardson PE (1984) Development of immunity to bacterial blight of cotton and its implications for other diseases. Plant Disease 68: 168-173.
- Bird LS (1982) The MAR (multi-adversity resistance) system for genetic improvement of cotton. Plant Disease 66: 172-176.
- Essenberg M, Bayles MB, Samad RA, Hall JA, Brinkerhoff LA, et al. (2002) Four near-isogenic lines of cotton with different genes for bacterial blight resistance. Phytopathology 92: 1323-1328.
- Koczan J, Albers D, Gholston K (2017) Identification of an alternative source of inoculum causing bacterial blight. Beltwide Cotton Conference Dallas, TX.






























