Isolation,Identification and Characterization Effective Rhizobium Species Nodulating Mung Bean (Vigna radiata) from some places of North Shewa
Asnake Beshah* and Fassil Assefa
Addis Ababa University, Ethiopia
Submission: March 07, 2019; Published: June 11, 20199
*Corresponding author: Asnake Beshah, Addis Ababa University, Addis Ababa, Ethiopia
How to cite this article: Asnake Beshah, Fassil Assefa. Isolation,Identification and Characterization Effective Rhizobium Species Nodulating Mung Bean (Vigna radiata) from some places of North Shewa. Int J Environ Sci Nat Res. 2019; 20(1): 5556026. DOI:10.19080/IJESNR.2019.20.556026
Abstract
Nineteen Mung bean root nodules were collected from farmer’s plots in north shoa zone of Kewet and Ensaro woreda. Three of the isolates were rejected in the process of isolation and presumptive tests. The rest were re -inoculated into their host for authentication. From the re-inoculated isolates, fifteen formed nodules in their host but one isolate failed to form nodule upon re-inoculation. The authenticated rhizobia were characterized based on their, morphological and eco-physiological features. From the preliminary screening 87% of the isolates were slow growing rhizobia. Symbiotic effectiveness of isolates showed significant difference in percent of effectiveness (34-92%) on the host plant. AAUMR2, AAUMR1, AAUMR3 and AAUMR7 were found to be the most effective isolates with percent effectiveness of 94%, 86%, 84%and 81% respectively. Most isolates were categorized under effective rate which possess percent of effectiveness 50-80%. Isolates were also tested for their tolerance to different eco-physiological features such as pH, temperature, salt concentration, intrinsic antibiotic resistance and utilization of different nitrogen sources. These tests showed the presence of wide physiological diversity between isolates. Plants inoculated with different isolates displayed differences in respect to nodule number, nodule dry weight and shoot dry weight. The numerical analysis of isolates based on forty-six phenotypic features showed the presence of diversity among isolates and statistical analysis indicates their diversity towards symbiotic effectiveness. Some isolates as AAUMR1 (Kewet), AAUMR3, and AAUMR7 (Ensaro) that are highly effective and tolerant to a wide range of phenotypic features are promising in the development of inoculant.
Keywords: Biological nitrogen fixation; Presumptive tests; Inoculants; Phenotypic diversity
Introduction
Mung bean (Vigna radiata) is a warm season annual grain legume. The optimum temperature range for good production is 27-30°C (Imrie, 1998). Mung bean requires 60-75 days to mature. It is useful crop in drier areas and has a good potential for crop rotation and relay cropping with cereals using residual moisture. The mung bean was also known as green gram or golden gram and is mainly cultivated in the Indian subcontinent. Now days; it is being cultivated after harvesting of Rabi crops (wheat, mustard, lentil, etc.). It can fit in as a cash crop between major cropping seasons. It is grown three times in a year covering 43,680ha with an average yield of 0.78t/ha. Besides being one of the shortest duration field crops in the world (can be harvested within two months), soil rhizobium bacteria around the mung bean root zone can symbiotically fix N2 gas from the air and this makes it among the most popular components in cropping systems. However, seed yield in farmers’ fields is still low, varying from 0.3 to 2.1t/ha [1].
Mung bean is one of the major calories (347-k cal. food energy) and protein (19.5% to 28.5%) sources in Asia, especially for the vegetarian population. High lysine content, which makes mung bean a good complementary food for rice-based diets, is usually the first limiting amino acid (Chen et al.1987). It is a popular food among vegetarians since it contains a lot of proteins and fiber and the main advantage of mung bean is that it helps in digestion and controls the amount of cholesterol content in our body. Mung bean contains a lot of minerals like calcium and potassium which was essential for enhancing the strength of bones and teeth. Fat content in mung bean is very low so it is highly recommended for people who want to minimize fat from their body. It is a store house of nutrients and it is a nutrition giving food and they are rich in Vitamin B, vitamin C, Manganese and a lot of other essential nutrients required for effective functioning of the human health (AVRDC, 1988).
In Ethiopia these crops also growing in smallholder farmers in drier marginal environments. As compared to other pulse crops, its production is around 8qt per hectare (SARC, 2005). However, for resource poor farmers in drier marginal environments it has been an important grain legume. These farmers need a variety and fertilizer that gives maximum production and stable yield in their environments. Several works showed that most of the rhizobia nodulation of legumes crop are very effective in soil fertility and nitrogen fixation [2,3]. However, the work on mung bean is very limited about isolation of rhizobia that nodulate mung bean.
This study is therefore focused on isolation and characterization of root nodule bacteria of Mung bean from one of the best pulses growing area of Amhara region of North shoa zones. The search for symbiotic effective isolates could contribute to the development of rhizobial inoculants to fully realize the potential of BNF in low input agriculture in the country.
Materials and Methods
Sampling sites
The selected sample sites of this study covered were in the major Mung bean growing areas of North shoa zones of the Amhara Region. In these areas Mung bean has been growing for a long time without any history of inoculation with rhizobia. The root nodules were collected between October and November 2013.
Sample collection and isolation of rhizobia
Collection of nodules
Nodule samples were randomly collected from the farmer’s field and immediately kept in sealed vials containing a desiccant (Silica gel) covered with 1cm of cotton wool for isolation of rhizobia [4].
Isolation of root nodule bacteria
Dehydrated or desiccated root nodules were immersed in sterile distilled water over night in labeled sets of flasks to imbibe water. The imbibed nodules were surface sterilized with 70% ethanol for 10 seconds and then to 3% (v/v) solution of sodium hypochlorite (NaHClO3) for 3 minutes according to Somasegaren & Hoben [4]. The surface sterilized nodules were then rinsed in five changes of sterile distilled water to completely rinse off and remove the sterilizing chemicals.
The surface-sterilized nodules were then transferred to sterile Petri-dishes and crushed with alcohol flamed sterile glass rod in a drop of normal saline solution (0.85% NaCl) inside a laminar flow hood. Stock solution of Congo red (CR) was prepared by dissolving 0. 25g of CR in 100ml sterile distilled water [5] and then 10ml of the CR stock solution was added to one liter of YEMA before autoclaving. Finally, Loopful of crushed nodule suspensions were streaked on YEMA plates with CR and incubated at 28 ± 20C for 3-7 days.
Purification and preservation of isolates
Single dome-shaped colonies were picked with sterile inoculating loop and streaked on sterile YEMA plates and incubated at 28+20C. The purity and uniformity of colony types were carefully examined through repeated re-streaking and a single well isolated colony was picked and transferred to YEMA slant containing 0.3% (W/V) CaCO3 in a culture tube and incubated at 28+20c. When enough growth was observed, the culture was transferred to be preserved at 40c for future use [5].
Presumptive test of the isolates
Each isolate was examined for presumptive purity using Peptone Glucose Test (PGT), gram staining and growth response to YEMA-CR medium [4].
Congo red absorption
Stock solution of Congo red was prepared by dissolving 0.25g of Congo red in 100ml of sterile distilled water. From stock solution, 10ml was added to a liter of YEMA and autoclaved. Loop full of test isolates were streaked on the medium and covered with aluminum foil to dark incubate at 28+20C for 3 to 7 days to detect Congo red absorption by the colonies [5].
Peptone glucose test
Peptone Glucose Test was prepared according to the procedure of Lupwayi & Haque [6] by dissolving 5g of glucose, 10g of peptone, 15g of agar and 10ml of bromo cresol purple (BCP) in a liter of distilled water and the pH was adjusted to 7.0 with 1N NaoH and acetic acid. Seven days old yeast extracts mannitol broth culture containing approximate number of cells (104 cells ml-1) was streaked on to the Peptone Glucose Medium to observe the growth after having incubated at 28+20C for 3 to 7 days [6].
Authentication of the isolates and preliminary screening of Symbiotic effectiveness of isolates on sand pot experiment
In order to test the definitive purity of all rhizobial isolates, nodulation test was carried out for each of the purified isolates. They were inoculated into the host plant potted in to 3Kg capacity plastic pots containing sterilized and nitrogen free sand [4]. The sand was thoroughly washed with 1N sulfuric acid whereas the pots were surface sterilized with 95% ethanol. Six seeds were sown in each pot and thinned down to three after germination. Each isolate was inoculated into Erlenmeyer flask and stay for 7 days and 1ml of the culture suspension was inoculated into each seedling [4].
Cultural characterization of isolates
Colony morphology
The morphological characteristics of the isolates were determined according to Lupwayi and Haque (1994). A loopful of old grown broth culture from each isolate was inoculated onto YEMA and incubated at 28±20C for 3-7days. After 7days, colony diameter, morphology and colony texture were recorded.
Acid-base production
To determine the ability of the rhizobial isolates to produce acid or alkaline in the medium, YEMA containing bromothymol blue (BTB) (0.025w/v) was used. A loop full of the isolates from a five days old culture broth was streaked on to the YEMA BTB medium and incubated for 3-7 days to record the color changes of the medium [7].
Biochemical and physiological tests
For each biochemical and physiological test, inoculation of a loopful of five days old broth culture was streaked on to the YEMA medium. The inoculated YEMA plates were incubated at 28 ± 20C for 3-5 days [4]. For each experiment, three replicates and controls were used per test as indicated in Maatallah et al. [8]. Ultimately, the growth of each rhizobial isolate was determined as (+) for positive growth, (++) for abundant growth and (-) for no growth.
Amino acid utilization
Different types of amino acids including L-arginine, L-glutamate, L-leucine, L-phenylalanine, L-tryptophan, urea and L-tyrosine were used in this experiment in order to determine the ability of the isolates to utilize the amino acids as a nitrogen source. These amino acids were added at a concentration of 0.5g/l to a basal media source that lack ammonium sulfate and supplemented with 1g/l of mannitol. The membrane filter sterilized amino acids were added to the autoclaved and cooled (approximately 550C) basal media as indicated in Amargar et al. [9]. Finally, five days rhizobial suspensions were inoculated into these basal media and incubated at 28±20 C for 3-5 days.
pH tolerance
The capacity of each rhizobial isolate to grow on acidic and alkaline media was determined by inoculating each isolate on YEMA adjusted at a pH of 4.0, 4.5, 5.0, 6, 7, 8, 8.5, 9.0, using 1N NaOH and acetic acid as described by Bernal & Graham [10].
Salt tolerance
The ability of the isolates to grow at different level of salt concentrations was determined by inoculating each isolate on the YEMA media containing 1%, 4%, 5%, 6%, 7%, 8%, 9% and 10% of NaCl as indicated in Lupwayi & Haque [6].
Temperature tolerance
The growth of each isolate at different incubation temperatures was evaluated by inoculating each isolate on YEMA plates. The inoculated plates were incubated at a temperature of 40C, 100C, 150C, 200C, 250C, 300C, 350C, 400C, 450C and 480C as indicated in Lupwayi & Haque [6].
Intrinsic antibiotic resistance
The resistance of isolates to different antibiotics at different concentration was evaluated by streaking each isolate on YEMA containing freshly prepared filter sterilized antibiotics using 0.22m sized membrane filters. The stock solution of each antibiotic was first prepared as described in Lupwayi & Haque [6] and was kept in refrigerator until they were used in the test. The antibiotics were Tetracycline, Erythromycin, Ampicillin, Chloroamphinicol, and penicillin. Each antibiotic was tested at the following concentrations. Ampicilin at 10g/ ml, Chloroamphinicol at 2. g/ml and 5g/ml, Tetracyclin at 2.5g/ ml, Erythromycin at (2.5g/ml, 5g/ml and 10g/ml) and penicillin at (2.5g/ml, 5g/ml and10 g/ml). Erythromycin was dissolved in ethanol, whereas the other four were dissolved in sterilized water. The stock solution of each antibiotic was prepared by dissolving 2g of each antibiotic in 100ml of water. The required concentration was aseptically added to the media using a single pipette for each antibiotic. The stock solution of each antibiotic was filter sterilized using a milli pore filter (0.22m) and aseptically added to autoclaved YEMA (kept at 500C in water bath) at the final concentrations of 2.5, 5 and 10g/ml, which is 12.5, 25 and 50l of antibiotic solution per 100ml medium, respectively, and finally poured separately in to plates.
Relative effectiveness of the isolates
After sixty days of planting upon re-inoculation, the plants were uprooted to measure nodule number, nodule dry weight and shoot dry weight. The effectiveness of isolates in accumulating plant shoot dry matter was calculated as described in Somasegaren & Hoben [4] and Molungoy [11] as follows:
SE = Inoculated plant D.M. X 100,
N-Fertilized plant D.M
Where, D.M. = dry matter, S.E. = symbiotic effectiveness
The rate of nitrogen fixing effectiveness is evaluated as: Highly effective > 80%, Effective 50- 80%, Lowly effective 35- 50% and Ineffective <35%.
Data analysis
Symbiotic effectiveness of the strains was measured in terms of the number of nodules, shoot dry weight and nodule dry weight from greenhouse trial. Phenotypic variability was analyzed using a computer cluster analysis applying the unweighted pair group method with the average (UPGMA) by PCORD statistical software ver. 5.0 of hierarchical clustering method [8]. One-way analysis of variance of data was also undertaken using the SPSS statistical program ver.17.0. Mean separation was calculated using the Tukey s values when the F-test was significant at P=0.05 [12-15].
Results and Discussion
Isolation and authentication of rhizobia
A total of 19 nodule samples were collected, 9 isolates from Ensaro and 10 isolates from Kewet werda. Three of them were rejected in the presumptive test and one is discarded upon authentication. All isolates did not grow on PGA and did not absorb congored on YEMA-CR media. Twelve of them changed YEMA-BTB medium into blue color whereas three of them changed into yellow. All but one isolate, formed nodules and authenticated as root nodule bacteria after they were reinoculated into the host plant [16-20].
Characterization of the isolates
Morphological and growth characteristics of isolates
Isolates were grown on YEMA medium to determine colony type, colony diameter and colony texture. About colony texture 46% showed large watery colonies (LW) and 53% of the isolates were characterized as large mucoid (LM) texture on YEMA media. The colony diameter of all the isolates ranged between 2mm and 4mm the largest colony diameter of 4 was observed on isolates AAUMR2 (Kewet) and AAUMR1 (Kewet) whereas the smallest diameter of 1 mm was recorded for isolates AAUMR9, (Ensaro).
Utilization of carbon sources
All the isolates were able to catabolize a large variety of carbon sources. All the isolates (100%) were found to catabolize dextrose, fructose, maltose, and sucrose. However, lactose, and cellulose were utilized by 86%. Isolates AAUMR3 and, AAUMR9 of ensaro, failed to grow on lactose whereas; all the rest grew very well. Eleven isolates (73% of the isolates) were found to catabolize and grew on the basal medium containing all the 6 tested carbon sources while isolates AAUMR1, AAUMR2, AAUMR13 and AAUMR14 of Kewet showed abundant growth on all tested carbon sources. On the other hand, the isolates AAUMR3 of Ensaro relatively utilized a smaller number of carbon sources (66%) out of the tested carbon sources [21-33] (Figure 1).

Amino acid utilization
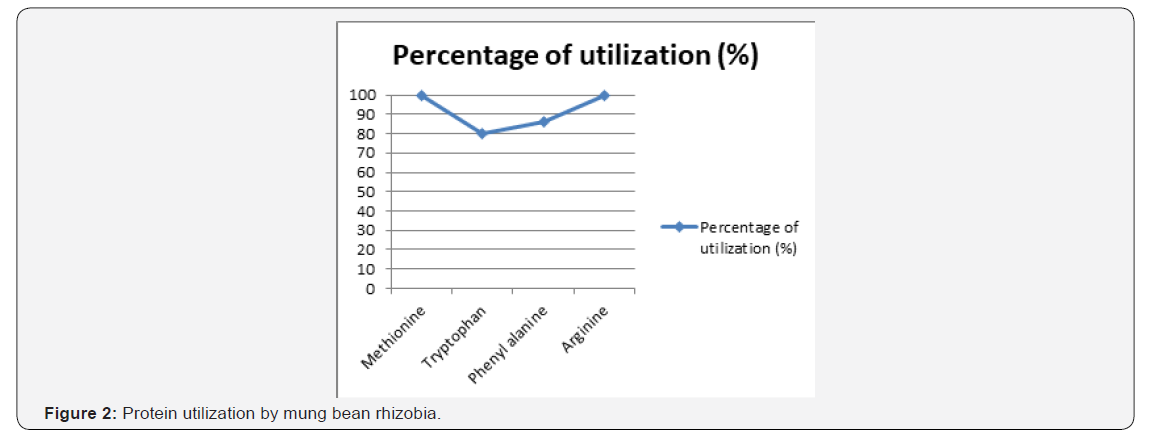
Almost all the isolates were able to catabolize a large variety of nitrogen sources. All the isolates (100%) utilized methionine and arginine likewise, 80 and 86%of the rhizobial isolates catabolized phenyl alanine and tryptophan, respectively. This is like the findings of Shraddha et al. (2013) on mungbean rhizobia isolated from different parts of India. Seventy three (73%) of the isolates that include AAUMR (4, 2, 5, 6, 13, 12, 14, and 10) of kewet and isolates AAUMR (8, 11, and 15) from Ensaro utilized 100% of the tested amino acids whereas, isolates AAUMR3 of Ensaro were found to be much fastidious with the ability of catabolizing 50% of the tested amino acids [34-40] (Figure 2).
pH tolerance
All isolates tolerated pH levels of 6-8 and 26% of the isolates were found to be tolerant to pH 4. The other 40% of the isolates grew at a pH 9. Isolates AAUMR3 and AAUMR13 from Kewet showed growth on all tested pH levels 4-9. This is in contrary to the reports of Shraddha et al (2013) that showed mung bean rhizobia grew on YEMA medium with pH levels of 5 - 8 It was also interesting to note that 13% of the isolates from Kewet and Ensaro tolerated wide range of pH 4-9 as opposed to the report of Shraddha et al (2013) who reported that mungbean rhizobia showed a neutral -tolerant tendency and also reported that 90% to 100% of the isolates from India grew on medium with lightly acidic (pH. 5) and neutral pH [41-95] (Figure 3).

Salt tolerance
Isolates displayed differences in growth on YEMA medium adjusted at different NaCl concentrations. All isolates were tolerant to salt concentration of 1% but showed a steady decrease in growth when they were inoculated into the medium containing 4 to 10% salt concentration. Consequently, 66% of the isolates grew at 4% NaCl, whereas 20% of isolates were resistant to salt concentration of 7%. Isolates AAUMR6 of Kewet were found to be the most tolerant strains that grew at Salt concentration of 9%. This is like the findings of Kucuk & Kivank [96] that salt tolerance by mungbean rhizobia from India. Isolates like AAUMR6 (Kewet) were the most tolerant to all salt concentrations the most sensitive isolates were AAUMR2, AAUMR5, and AAUMR12 (Kewet) and AAUMR15 (Ensaro), that were able to grow at salt concentration of only 1% (Figure 4).

Temperature tolerance
All isolates were able to grow within temperature range of 20 to 350c. Isolates AAUMR4, AAUMR2, AAUMR1, AAUMR10, and AAUMR14 (Kewet) and AAUMR3 AAUMR9, AAUMR8 and AAUMR15 (Ensaro) were grown at the lowest temperature of 40C. There was a progressive decrease in growth of isolates from a temperature range of 400C to 450C. Only isolates AAUMR2, AAUMR10, and AAUMR14 (Kewet) and AAUMR9 and AAUMR15 (Ensaro) were grown at all tested incubation temperature (4-450C) (Figure 5).

Intrinsic antibiotic resistance
All the isolates (100%) were found to be resistant to all the tested antibiotics at a concentration of 2.5mg/ml (Table 7). Isolate AAUMR9 (Ensaro) was found to be the most resistant of the isolates to different antibiotics followed by isolate AAUMR6 and AAUMR12 (Kewet) .The most sensitive isolate was AAUMR6; did not grow on YEMA medium containing any antibiotic at concentration of 5 and 10g/ml, followed by isolates AAUMR4, AAUMR10 (Kewet) and AAUMR15 (Ensaro) (Figure 6).

Conclusion
The result of this work is an indicative of the presence of wide diversity in rhizobial isolates of Mungbean collected from different growing areas of North shoa zone of the Amhara Regional State. Diversity of the isolates was observed based on their morphological, physiological, host infection and symbiotic effectiveness. This study showed that rhizobial isolates showed diversity with respect to symbiotic infectiveness and effectiveness with their host. Similarly, the numerical analysis also confirmed that these isolates were phenotypically diverse which indicates that their real diversity must also be confirmed by genetic analysis using molecular techniques.
The tolerance of isolates to different pH levels, temperature, salinity, and antibiotics is an important quality of rhizobial strains to screen and develop inoculants that are endowed with ecological competitiveness. In the presence of different environmental stresses, the tolerant isolates would survive, occupy nodules, and fix nitrogen and provide the host to boost crop production. Accordingly, the following isolates were found to be effective in nitrogen fixation and resistant to different environmental stresses. In most of the tests, the highly effective AAUMR2, AAUMR3, AAUMR1 and AAUMR7 was found to be the best of all the isolates followed by isolates AAUMR4, AAUMR5, AAUMR6, AAUMR13, AAUCR12, AAUMR10, AAUMR8 and AAUMR14.
References
- Herridge D, Rose I (2000) Breeding for enhanced nitrogen fixation in crop legumes. Field crops res 65(2-3): 229-248.
- Tesfaye G (2008) Symbiotic and phenotypic diversity of rhizobial isolatesnodulating Vicia faba from western Shoa and Hararghe, Ethiopia. M.SC. Thesis, Addis Ababa University, Ethiopia.
- Belay Z (2006) Symbiotic and Phenotypic Diversity of Rhizobium leguminosarum var viceae Isolates (Vicia faba) from Northern Gondar, Ethiopia. M.sc. thesis. Addis Ababa University, Ethiopia. pp. 1-73.
- Somasegaren P, Hoben HJ (1994) Handbook for rhizobia. Methods in legume-Rhizobium technology. Springer verlag, New York, USA, pp. 1-441.
- Vincent JM (1970) A Manual for the Practical Study of Root Nodule Bacteria. Blackwell, Oxford and Edinburgh, p. 164.
- Lupwayi NZ, Haque I (1994) Working document: legume-rhizobium technology manual. Environmental science division international livestock center for Africa, Addis Ababa Ethiopia, pp. 1-40.
- Jordan DC (1984) Family III. Rhizobiaceae. In: Krieg NR, Holt JG (Eds.), Bergey s Manual of Systematic Bacteriology. The Williams and Wilkins, Baltimore, pp. 234-254.
- Maatallah J, Berraho EB, Sanjuan J, Lunch C (2002) Phenotypic characterization of rhizobia isolates from chick pea (Cicer arrientium) growing in Morroccan soils. Agronomic 22: 321-329.
- Amargaer N, Macheret V, Aguerre G (1997) Rhizobum gallicum sp. Nov, and Rhizobum giardinii sp. Nov. from Phaseolus vulgaris nodules. Int J syst Bacteriol 47(4): 996-1006.
- Bernal G, Graham PH (2001) Diversity in the rhizobia associated with Phaseolus vulgaris L. in Ecuador, and comparisons with Mexican bean rhizobia. Canadian J Microbiol 47(6): 519-526.
- Mulongoy K (2004) Technical paper 2: Biological nitrogen fixation. pp. 1-3.
- Alexander M (1997) Introduction to soil microbiology. (2nd edn), John Willy and Sons, Inc, USA, pp. 467.
- Arayankoon T, Schomberg HH, Weaver RW (1990) Nodulations and N2 fixation of guar at high temperature. Plant and Soil 126(2): 209-213.
- Atkins CA (2004) Efficiencies and inefficiencies in legume / Rhizobum symbiosis- A review. Plant and Soil 82(3): 273-284.
- Badaruddin M, Meyer DW (1994) Grain legume effects on soil nitrogen grain yield and nitrogen nutrition of wheat. Crop Sci 34(5): 1104-1309.
- Baginsky C, Brito B, Imperial J, Palacios JM, Ruiz-Argüeso T (2002) Diversity and evolution of hydrogenase systems in rhizobia. Applied Environ Microbiol 68(10): 4915-4924.
- Beijerinck MW (1888) Rhizobial systematic. Bot Ztg 46: 796-804.
- Benson DR, Clawson ML (2000) Evolution of the actinorhizal plant symbioses. In: Prokaryotic Nitrogen Fixation: A Model System for Analysis of Biological Process.
- Bernstein L, Ogata G (1966) Effects of salinity on nodulation, nitrogen fixation and growth of soybeans and alfalfa. Agron J 58(2): 203-210.
- Bordeleau LM, Prevost D (1994) Nodulation and nitrogen fixation in extreme environments. Plant and Soil 161(1): 115-124.
- Boddey RM, Peoples MB, Palmer B, Dart PJ (2000) Use of the 15N natural abundance method to quantify biological nitrogen fixation in woody perennials. Nutrient Cycling in Agro-ecosystems 57(3): 235- 270.
- Brockwell J, Bottomley PJ, Thies JE (1995) Manipulation of rhizobia microflora for improving crop productivity and soil fertility. Plant Soil 174(1/2): 143-180.
- Broughton WJ, Samery U, Stanley J (1987) Ecological genetics of Rhizobummeliloti: symbiotic plasmid transfer in the Medicago sativa rhizosphere. FEMS Microbiol Letters 40(2-3): 251-255.
- Brencic A, Winans SC (2005) Detection of and response to signals involved in host microbe interactions by plant-associated bacteria. Microbiol Mol Biol Rev 69(1): 155-194.
- Chen WM, Moulin L, Bontemps C, Vandamme P, Béna G, et al. (2003) Legume symbiotic nitrogen fixation by ß-proteobacteria is widespread in nature. J Bacteriol 185(24): 7266-7272.
- Chen WM, Laevens S, Lee TM, Coenye T, de Vos P, et al. (2001) Ralstonia taiwanensis sp. nov. isolated from root nodules of Mimosa species and sputum of a cystic fibrosis patient. Int J System and Evol Microbiol 51(Pt 5): 1729- 1735.
- Cordovilla MP, Ocana A, Ligero F, Lluch C (1995) Salinity effects on growth analysis and nutrient composition in four grain legumes Rhizobium symbiosis. J plant Nutr 18: 1595-1609.
- Cordovilla MP, Ligero F, Lluch C (1994) The effect of salinity on N fixation and assimilation in Vicia faba. J Exp Bot 45(10): 1483-1488.
- Correa OS, Barneix AJ (1997) Cellular mechanisms of pH tolerance in Rhizobium loti. World J Microbiol Biotechnol 13(2): 153-157.
- Crews TE, Peoples MB (2004) Legume versus fertilizer sources of nitrogen ecological trade off and human needs. Agric Econ Environ 102: 279-297.
- Csonka LN, Hanson AD (1991) Prokaryotic Osmoregulation: genetics and physiology. Annu Rev Microbiol 45: 569-606.
- De-Faria SM, Lewis GP, Sperennt JI, Southerland JM (1989) Occurrence of nodulation on Leguminosae. New phytol 111(4): 607-609.
- Delgado MJ, Ligero F, Lluch C (1994) Effects of salt stress on growth and nitrogen fixation by pea, faba-bean, common bean and soybean plants. Soil Biol Biochem 26(3): 371-376.
- Dilworth MJ, Howieson JG, Reeve WG, Tiwari RT, Glenn AR (2001) Acid tolerance in legume root nodule bacteria and selecting for it. Aust J Exp Agric 41(3): 446-453.
- Dowling DN, Broughton WJ (1986) Competition for nodulation of legumes. Ann Rev Microbiol 40: 131-157.
- Frank B (1889) Rhizobial systematic. Beru Devt Bot Ges 7: 322-346.
- Freiberge C, Rosenthal A, Fellay RW, Broughton J, Perret X, et al. (1997) Molecular basis of symbiosis between rhizobium and leumes. Nature 387(6631): 394-401.
- Fuhrmann J, Davey CB, Wollum AG (1986) Desiccation tolerance in clover rhizobia in sterile soils. Soil Sci Soc Am J 50: 639-644.
- Gage DJ (2004) Infections and invasion of roots by symbiotic, nitrogenfixing rhizobia during nodulations of temperate legumes. Microbiol Mol Biol Rev 68(2): 280-300.
- Giller KE (2003) Biological nitrogen fixation. In: Schroth G, Sinclair FL (Eds.), Trees, crops, and soil fertility. CABI publishing, Walling Ford, UK, pp. 259-270.
- Giller KE (2001) Nitrogen Fixation in Tropical Cropping systems. (2nd edn), CABI Publishing, Walling Ford, UK, p. 448.
- Graham PH, Dreeger KJ, Ferrey ML, Conroy MJ, Hammes BE, et al. (1994) Acid pH tolerance in strains of Rhizobium and bradyrhizobium, and initial studies on the basis for acid tolerance of Rhizobium tropic UMR1899. Can J Microbiol 40: 198-207.
- Graham PH (1992) Stress tolerance in Rhizobium and bradyrhizobium and nodulation under adverse soil conditions. Can J Microbiol 38(6): 475-484.
- Guerin V, Trinchant JC, Rigaud J (1991) Nitrogen fixation (C2H2) reduction by broad bean (Vicia faba L.) nodules and bacteroids under water restricted conditions. Plant Physiol 92(3): 595-601.
- Hill S (1992) Physiology of nitrogen fixation in free-living heterotrophs. In: Stacey G, Burris RH, Evans HJ (Eds.), Biological nitrogen fixation. New York, Chapman & Hall, pp. 87-134.
- Howieson JG, Hara GWO, Carr SJ (2000). Changing roles of legumes in Mediterranean agriculture, developments from an Australian perspective. Field Crops Research 65(2-3): 107-122.
- Hubbell DH, Kidder G (2003) Biological nitrogen fixation. Food and agricultural Sci 16: 1-4.
- Hungaria M, Vergas MAT (2000) Environmental factors affecting N2 fixation in grain legumes in tropics, with an emphasis on Brazil. Field crops research 65: 151-164.
- Hussain N, Mujeeb F, Tahir M, Khan GD, Hassan NM, et al. (2002) Effectiveness of Rhizobium under salinity stress. Asian J Plant Sci 1(1): 12-14.
- http://edis.ifas.ufl.edu/ss180
- http://www.biology.ed.ac.uk/research/groups/jdeacon/microbes/ nitrogen
- Jarabo-Lorenzo A, Perez-Galdona R, Donate-Correa J, Rivas R, Velazquez E, et al. (2003) Genetic diversity of bradyrhizobial populations from diverse geographic origins that nodulate Lupinus spp. and Ornithopus spp. Syst Appl Microbiol 26(4): 611-623.
- Jordan DC (1982) Transfer of Rhizobium japonicum Buchanan 1980 to Bradyrhizobium gen. nov., a genus of slow-growing, root nodule bacteria from leguminous plants. Int J Syst Evol Microbiol 32: 136-139.
- Kishinevsky BD, Sen D, Weaver RW (1992) Effect of high root temperatures on Bradyrhizobium peanut symbiosis. Plant and Soil 143(2): 275-282.
- Kouchi H, Akao S, Yoneyama T (1986) Respiratory utilization of Clabeled photosynthate in nodulated root systems of soybean plants. J Exper Bot 37(7): 987-993.
- Lapinskas E, Ambrazaitiene D, Piaulokaite-motuziene L (2005) Estimation of soil microbiological properties in relation to soil acidity and fertilization. Latvian J Agro 8: 39-43.
- Latta RA, Carter ED (1998) Increasing production of an annual medic – wheat rotation by grazing and grass removal with herbicides in the Victorian mallee. Aust J Exp Agr 36(3): 211-217.
- Lodwig EM, Hosie AHF, Bourde´s A, Findlay K, Allaway D, et al. (2003) Amino-acid cycling drives nitrogen fixation in legume rhizobium symbiosis. Nature 422(6933): 722-726.
- Menna P, Hungria M, Barcellos FG, Bangel EV, Hess PN, et al. (2006) Molecular phylogeny based on the 16S rRNA gene of elite rhizobial strains used in Brazilian commercial inoculants. System Appl Microbiol 29(4): 315-332.
- Michelle IL (2007) Diversity of root nodule bacteria associated with Phaseolus coccineus and Phaseolus vulgaris species in South Africa. M. SC. Thesis. University of Pretoria, South Africa. pp. 1-130.
- Michiels J, Verreth C, Vanderleyden J (1994) Effects of temperature stress on bean nodulating Rhizobium strains. Appl Environ Microbiol 60(4): 1206-1212.
- Moawad H, Beck D (1991) Some characteristics of Rhizobium leguminosarum isolate from uninoculated field-grown lentil. Soil Biol Biochem 23(10): 917-925.
- Moulin L, Munive A, Dreyfus B, Boivin-Masson C (2001) Nodulation of legumes by members of the beta-subclass of Proteobacteria. Nature 411(6840): 948-950.
- Norris DO (1965) Legume-rhizobium association. Plant soil 22: 143- 146.
- Nour SM, Cleyet Marel J, Beck D, Effosse A, Fernandez MP (1994) Genotypic and phenotypic diversity of rhizobium isolated from chickpea (Cicer ariethinum L.). Can J microbial 40(5): 345-354.
- Now SM, Fernandez MP, Normand P, Cleyet-Marel JC (1994) Rhizohiurn ciceri sp. nov., consisting of strains that nodulate chickpeas (Cicer arietinum L.). Int J Syst Bacteriol 44(3): 511-522.
- Peoples MB, Herridge DF, Ladha JK (1995) Biological nitrogen fixation: an efficient source of nitrogen for sustainable agricultural production. Plant Soil 174(1-2): 3-28.
- Prell J, Poole P (2006) Metabolic changes of rhizobia in legume nodules. Trends in Microbiol 14(4): 161-168.
- Rome S, Fernandez MP, Brunel B, Normand P, Cleyet-Marel JC (1996) Sinorhizobium medcae spp. nov. isolated from annual Medicago spp. Int J Syst Bacteriol 46(4): 972-980.
- Roughley RJ (1970) The influence of root temperature, Rhizobium strain and host selection on the structure and nitrogen-fixing efficiency of the root nodules of Trifolium subterraneum. Ann Bot 34(3): 631-646.
- Rubio LM, Ludden PW (2005) Maturation of nitrogenase: a biochemical puzzle. J Bacteriol 186(2): 405-414.
- Smith LT, Allaith AM, Smith GM (1994) Mechanisms of osmotically regulated Nacetylglutaminylglutamine amide production in Rhizobium meliloti. Plant Soil 161: 103-108.
- Sprent JI, Sprent P (1990) Nitrogen fixing organisms. Pure and applied aspects. Chapman and Hall, London, United Kingdom.
- Sprent JI (1972) The effects of water stress on nitrogen fixing root nodules. IV. Effects on whole plants of Vicia faba and Glycine max. New Phytol 71(4): 603-611.
- Tate RL (1995) Soil microbiology (symbiotic nitrogen fixation), John Wiley and Sons, Inc, New York, USA, pp. 307-333.
- Talukder MS, Solaiman A, Khanam D, Rabbani MG (2008) Characterization of Some Rhizobium Isolates and Their Effectiveness on Pea. Bangladesh J Microbiol 25(1): 45-48.
- Thies JE, Singleton PW, Bohlool BB (1991) Influence of the size of indigenous rhizobial populations on the establishment and symbiotic performance of introduced rhizobia on field-grown legumes. Appl Environ Microbiol 57(1): 19-28.
- Thomas RJ (1995) Role of legumes in providing N for sustainable tropical pasture systems. Plant and Soil 174(1/2): 103-118.
- Vandamme P, Goris J, Chen WM, de Vos P, Willems A (2003) Burkholderia tuberum sp. nov. and Burkholderia phymatum sp.nov. nodulate the roots of tropical legumes. Syst Appl Microbiol 25(4): 507-512.
- Valverde A, Velázquez E, Fernandez-Santos F, Vizcaino N, Rivas R, et al. (2005) Phyllobacterium trifolii sp. nov., nodulating Trifolium and Lupinus in Spanish soils. J System Evol Microbiol 55(Pt 5): 1985-1989.
- Van Rossum D, Schuurmans FP, Gillis M, Muyotcha A, Van Verseveld HW, et al. (1995) Genetic and phenotypic analysis of Bradyrhizobium strains nodulating Peanut (Arachis hypogae L.) roots. Appl Environ Microbiol 61(4): 1599-1609.
- Vanparys B, Heylen K, Lebbe L, De Vos P (2005) Devosia limi sp. nov., isolated from a nitrifying inoculum. J System Evol Microbiol 55(Pt 5): 1997-2000.
- Versalovic J, Koeuth T, Lupski JR (1991) Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res 19(24): 6823-6831.
- Vlassak KM, Vandurleyden J (1997) Factors influencing nodule occupancy by inoculants rhizobia. Crit Rev Plant Sci 16: 163-229.
- Walsh KB (1995) Physiology of the legume nodule and its response to stress. Soil Biol Biochem 27(4-5): 637-655.
- Wani SP, Rupela OP, Lee KK (1995) Sustainable agriculture in the semiarid tropics through biological nitrogen fixation in grain legumes. Plant Soil 174(1-2): 29-49.
- White D (1995) The Physiology and Biochemistry of Prokaryotes. Oxford University Press, pp. 34-46.
- Willems A (2006) The taxonomy of rhizobia: an over review. Plant and Soil 287(1-2): 3-14.
- Wilson JK (1944) Over five hundred reasons for abandoning the crossinoculation groups of the legumes. Soil Sci 58: 61-69.
- Winch T (2006) Growing Food: A Guide to Food Production. Springer, Dordrecht, the Netherlands, pp. 150-151.
- Wood M, Cooper JE, Holding AJ (1984) Soil acidity factors and nodulation of Trifolium repens. Plant and Soil 78(3): 369-379.
- Young JPW, Haukka KE (1996) Diversity and phylogeny of rhizobia. New Phytol 133(1): 87-94.
- Zahran HH (1999) Rhizobium-Legume symbiosis and nitrogen fixation under severe conditions and in an arid climate. Microbiol Mol Biol Rev 63(4): 968-989.
- Zahran HH (1997) Diversity, adaptation and activity of the bacterial flora in saline environments. Biol Fertil Soils 25(3): 211-223.
- Zahran HH (1991) Conditions for successful Rhizobuim-legume symbiosis in saline environments. Biol Fertil Soils 12(1): 73-80.
- Küçük ÇD, K vanc M (2008) Preliminary characterization of Rhizobium strains isolated from chickpea nodules. Afri J Biotechnol 7: 772-775.






























