Abstract
Introduction: Ovarian cancer (OC) has a high mortality rate and limited treatments, necessitating novel therapeutic targets and biomarkers. Objective: Identify differentially expressed genes (DEGs) in OC, assess their prognostic value, and discover potential therapeutic molecules through gene expression analysis and computational docking. Materials and Methods: Analyzed 4204 DEGs using gene expression profiling. Survival and protein-protein interaction (PPI) networks identified key genes. GEPIA and Oncomine validated expression and prognosis. Virtual screening and Connectivity Map (CMap) analysis pinpointed inhibitors. Results: Among DEGs, 100 had survival significance. Key pathways included p53 signalling and cell cycle regulation. Eight hub genes (AURKB, CCNB1, CCNB2, CDK1, CENPA, CENPF, FOXM1 and NEK2) were central to disease progression. FOXM1, overexpressed in OC, was a promising target. ZINC13597767 was identified as a potential FOXM1 inhibitor. CMap linked trichostatin A and vorinostat to OC gene suppression. Discussion: Cell cycle genes, especially FOXM1, drive OC progression. Computational screening highlights FOXM1 inhibitors and epigenetic drugs (trichostatin A, vorinostat) as therapeutic candidates. Conclusion: FOXM1 is a key therapeutic target. Integrating transcriptomics and drug screening accelerates OC drug discovery, with FOXM1 inhibitors and epigenetic agents warranting further study.
Keywords: Differential Expression; FOXM1; Ovarian Cancer; Small Molecule Screening
Introduction
Ovarian cancer (OC) is one of the most formidable challenges in gynecological oncology owing to its high mortality rate, late-stage diagnosis, and resistance to conventional therapies [1]. Despite advancements in surgical techniques and chemotherapeutic regimens, the prognosis of patients with advanced-stage OC remains dismal [2], highlighting the urgent need for novel therapeutic strategies. The absence of effective early detection methods further complicates this problem [3], resulting in the majority of cases being diagnosed at a late stage when treatment options are limited and often ineffective [4]. Recent technological advancements in high-throughput genomics have revolutionized our understanding of cancer biology [5]. Gene expression profiling, enabled by tools such as microarrays and RNA sequencing, has provided deep insights into the molecular mechanisms underlyning OC [6]. Datasets such as those available from the Gene Expression Omnibus (GEO) have been pivotal in identifying differential gene expression profiles that shed light on OC pathogenesis [7]. For instance, comparative analyses of high-grade serous carcinoma and normal ovarian epithelium, such as those provided by datasets GSE7463 and GSE14407, have elucidated the key oncogenes and tumor suppressor genes that drive the aggressive nature of the disease [8]. To address the complexity of OC, we employed a multifaceted approach that integrated comprehensive gene expression analysis with functional validation to identify novel therapeutic targets. Differentially expressed genes (DEGs) were identified using GEO2R [9], adhering to stringent selection criteria (adjusted p < 0.05, and log2(fold change) > 1.5) to ensure the biological relevance of the findings. The use of Venn diagrams to visualize DEG overlaps across multiple datasets allows for the identification of consistent gene targets [10], providing a robust foundation for subsequent analyses.
To further elucidate the biological significance of these DEGs, functional annotation was performed using Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses using the DAVID database [11]. These analyses revealed the biological processes (BP), molecular functions (MF), and cellular components associated with the DEGs, as well as the pathways that they regulate. Such functional insights are crucial for identifying the molecular mechanisms underlying OC and potential pathways for therapeutic interventions. Additionally, a protein-protein interaction (PPI) network analysis was conducted using STRING and Cystoscope [12] to explore the interactions between DEGs and identify key regulatory nodes. This network-based approach highlights central genes and modules with significant interactions, which are prime candidates for therapeutic targeting. The identification of hub genes with high centrality within the network provided valuable insights into potential drug targets and their roles in OC pathology. To validate the clinical relevance of these identified hub genes, their expression levels were assessed using resources such as GEPIA and Oncomine [13]. These tools facilitate the comparison of gene expression between OC and normal tissues, and offer insights into the prognostic significance of these genes. Immunohistochemical analysis of patient tissue samples was also performed [14] to confirm gene expression patterns, link gene expression data with clinical outcomes, and enhance the translational relevance of the findings.
In parallel, an in silico screening approach was used to identify potential small molecule inhibitors. The Connectivity Map (CMap) database was utilized [15] to query dysregulated genes and identify compounds capable of reversing the OC gene expression signature. Molecular docking studies using Auto Dock Vina and Swiss Dock provided insights into the binding affinities of these compounds, highlighting promising candidates for further preclinical and clinical evaluation. By integrating gene expression profiling with functional annotation and small-molecule screening, this study aimed to identify novel therapeutic strategies for OC treatment. The combination of high-throughput data analysis with experimental validation and computational drug screening represents a comprehensive approach to address critical challenges in OC treatment. This integrative methodology not only enhances our understanding of the molecular mechanisms driving OC but also paves the way for developing targeted therapies that could significantly improve patient outcomes.
Methods
Data Acquisition
Two OC microarray datasets were retrieved from the GEO database GSE7463: HG_U95Av2 Affymetrix Human Genome U95 Version 2 Array, consisting of 43 high-grade serous OC samples and 10 normal ovarian surface epithelium samples [16]. GSE14407: Affymetrix Human Genome U133 Plus 2.0, which included 12 serous papillary OC samples and 12 normal ovarian epithelium samples.
Patient Characteristics and Clinical Data
The baseline characteristics of the patients included in the analysis were summarized, focusing on age, International Federation of Gynecology and Obstetrics (FIGO) stage, histological grade, CA125, and HE4 levels [17]. Associations between gene expression and clinical outcomes, including overall survival (OS) and progression-free survival (PFS), were analyzed using the Kaplan-Meier (K-M) and Cox proportional hazards models.
Data Preprocessing
Data preprocessing was performed using GEO2R to identify DEGs between the OC and normal samples. A cutoff threshold of adjusted p-value < 0.05 and |log2 fold change| > 1.5 was applied. The batch effects were corrected as necessary [18]. The Venn Diagram web tool (Venny 2.1) was used to determine the overlapping DEGs between the two datasets [19].
Functional Annotation and Pathway Enrichment
GO analysis classified DEGs into BPess, MF, and cellular component categories [20]. KEGG pathway enrichment was performed using the DAVID Bioinformatics Tool (v6.8) with a significance threshold set at p < 0.05.
Protein-Protein Interaction (PPI) Network Construction
The CMap database (http://www.broadinstitute.org/cmap/) was used to identify potential drugs for patients based on the gene signature of OC. Common dysregulated probe sets were queried using the CMap database. The upregulated and downregulated genes were converted to GPL571 probe IDs using Perl scripts and were entered into the CMap official online “rapid query” tool. The enrichment scores ranged from -1 to 1. A positive connectivity score (closer to 1) indicates a drug capable of inducing the OC cell signature, whereas a negative score (closer to -1) indicates a drug that can reverse it. Negative connectivity scores were confirmed as candidate molecules with potential therapeutic value, and details were retrieved from the PubChem database (https://pubchem. ncbi.nlm.nih.gov/).
Identification of Hub Genes
Hub genes were identified based on degree centrality in the PPI network. The top eight genes with the highest node degrees were selected as hub genes [21]. Validation was performed using The Cancer Genome Atlas (TCGA) data via GEPIA, comparing OC and normal tissues with |log2FC| > 1.5 and p < 0.05.
Validation of Hub Genes
Intersecting genes from the enrichment pathways and top eight nodes in the PPI network were identified as hub genes. The “Expression analysis Box Plots” module of GEPIA (http:// gepia.cancer-pku.cn) was utilized to analyze hub gene mRNA levels in OC, using default settings (|Log2FC| Cutoff: 1.5, p-value Cutoff: 0.05, and “Match TCGA normal and GTEx data”) [22]. The Oncomine database (https://www.oncomine.org/) was used to compare the expression levels in OC and normal specimens, with a fold change threshold of 1.5, p-value of 0.05, and gene ranking of 10%.
Immunohistochemical results of the hub genes were obtained from the Human Protein Atlas (version 18, https://www. proteinatlas.org/). The K-M plotter (http://kmplot.com/analysis) was used for survival analysis of the eight hub genes, with a total of 1,656 OC patients included in the OS analysis. The genes and their corresponding Affy IDs are as follows: CCNB1 (214710_s_at), AURKB (204092_s_at), CCNB2 (202705_at), CDK1 (203468_at), CENPA (204962_s_at), CENPF (207828_s_at), FOXM1 (202580_x_ at), RRM2 (202580_x_at), NEK2 (204641_at). The median expression of hub genes across all samples was chosen as a cutoff to categorize samples into high and low expression groups, followed by K-M analysis to assess patient prognosis.
FOXM1-Centric Protein-Protein Interaction Network and functional enrichment Analysis
A bioinformatics approach was used to analyze the FOXM1 PPI network in OC. FOXM1-associated proteins were identified using STRING, GO, and KEGG pathway enrichment analyses using DAVID and Metascape. Significant biological processes, cellular components, and MFs were identified with p-values of < 0.05. The results were visualized using Cystoscope to explore FOXM1’s role in cell cycle regulation and cancer progression, thereby highlighting its potential as a therapeutic target.
Small Molecule Identification
The CMap database (http://www.broadinstitute.org/cmap/) was used to identify potential drugs for OC patients based on the gene signature. Common dysregulated probe sets were used to query the CMap database [23]. Using Perl, upregulated and downregulated genes were converted to GPL571 probe IDs and entered into the CMap “rapid query” tool. Enrichment scores ranging from -1 to 1 were calculated, with negative scores confirmed as candidate therapeutic molecules, whose tomographs were further queried in the PubChem database [24].
In Silico Molecular Docking
In silico screening was conducted using the Auto Dock Vina 1.0.2. The protein structure was obtained from the Protein Databank (www.pdb.org) and prepared for docking using the Auto Dock tools. Water molecules were removed and charges and nonpolar hydrogens were added using MGL Tools [25].
Structural representations of the National Cancer Institute (NCI) diversity set II were obtained from the NCI website and converted to PDB format. Individual PDB files for docking were prepared using prepare_ligand.py scripts from MGL Tools 1.5.4, focusing on the largest non-bonded fragment present [26]. The docking grid size was adjusted to encompass the entire FOXM1 structure or tested derivatives. The docking stringency was set to eight, which is the default parameter. Swiss Dock was used as a secondary validation tool for docking, using the same protein structure (3G73) from the Protein Databank. Ligands were prepared using the Chimera program.
Binding Affinity Analysis
The docking results were analyzed by calculating the free energy of binding (kcal/mol), where lower values indicate stronger binding affinity. Swiss Dock served as a secondary validation tool for docking, and binding modes were evaluated based on Full Fitness and clustering. The results with the highest affinity scores were further assessed for potential therapeutic effects.
Software and Tools
All bioinformatic analyses were conducted using publicly available online platforms, including GEO2R, STRING, DAVID, GEPIA, Oncomine, and K-M Plotter. Molecular docking was performed using Auto Dock Vina and Swiss Dock, and Cyto scape was used for PPI visualization.
Statistical Analysis
Statistical analysis was conducted using SPSS (version 26.0). Chi-square tests were used to assess the associations between categorical variables, including the clinicopathological characteristics of patients with OC. Statistical significance was set at p < 0.05. These tools allow data quantification, visualization, and statistical comparisons to be performed efficiently across all experiments.
Results
A total of 4,204 DEGs were identified in OC samples, with 100 of these genes showing a significant association with OS in OC patients. The intersection between DEGs and OS-related genes revealed 16 key genes with differential expression and prognostic significance (Figure 1). Further analysis identified 432 overlapping genes (Table 1), consisting of 159 upregulated and 274 downregulated genes, derived by comparing gene expression profiles of OC with the GSE7463 and GSE14407 datasets. Venn diagram analysis demonstrated pathway-specific and shared gene enrichments across apoptosis, PI3K, MAPK, and cell cycle signaling, with MAPK uniquely contributing the largest subset (171 genes, 23.9%), underscoring its critical role in OC pathophysiology.
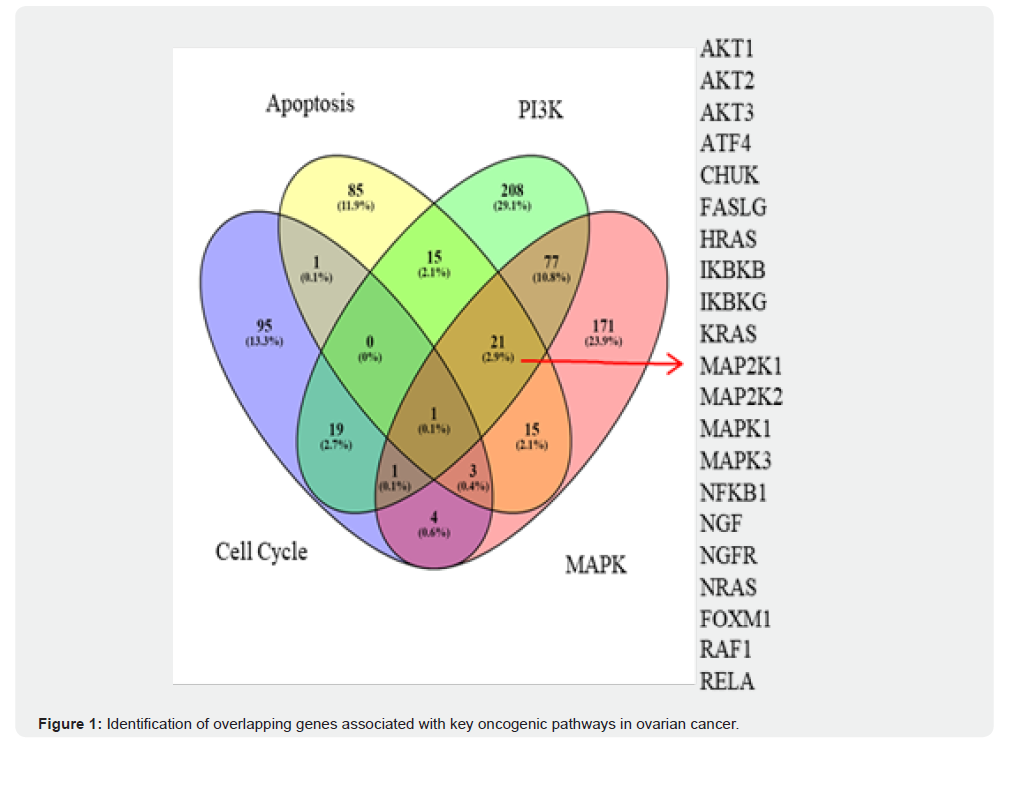

A total of 4,204 differentially expressed genes (DEGs) were identified in OC samples, among which 100 genes showed significant association with overall survival (OS) in OC patients. The intersection between DEGs and OS-related genes revealed 16 key genes with both differential expression and prognostic significance. Comparative analysis with GSE7463 and GSE14407 datasets identified 432 overlapping DEGs, including 159 upregulated and 274 downregulated genes. The Venn diagram illustrates the distribution of DEGs across four critical signalling pathways apoptosis, PI3K, MAPK, and cell cycle highlighting the genes uniquely enriched in each pathway as well as shared subsets across multiple pathways. These overlapping genes represent potential drivers of OC pathophysiology and candidate prognostic markers.
GO analysis of the shared DEGs was conducted using the ggplot2 R package, focusing on the top 10 significantly enriched GO terms and pathways based on p-values. GO BP analysis revealed that these DEGs were significantly enriched in processes associated with cell cycle regulation, particularly mitotic transitions and chromatid segregation. Notably, the highest fold enrichment was observed in pathways regulating the mitotic metaphase-anaphase transition (Figure 2A), indicating that dysregulation of these processes may be pivotal in OC pathology, as well as in the biological conditions reflected by the GSE7463 and GSE14407 datasets.
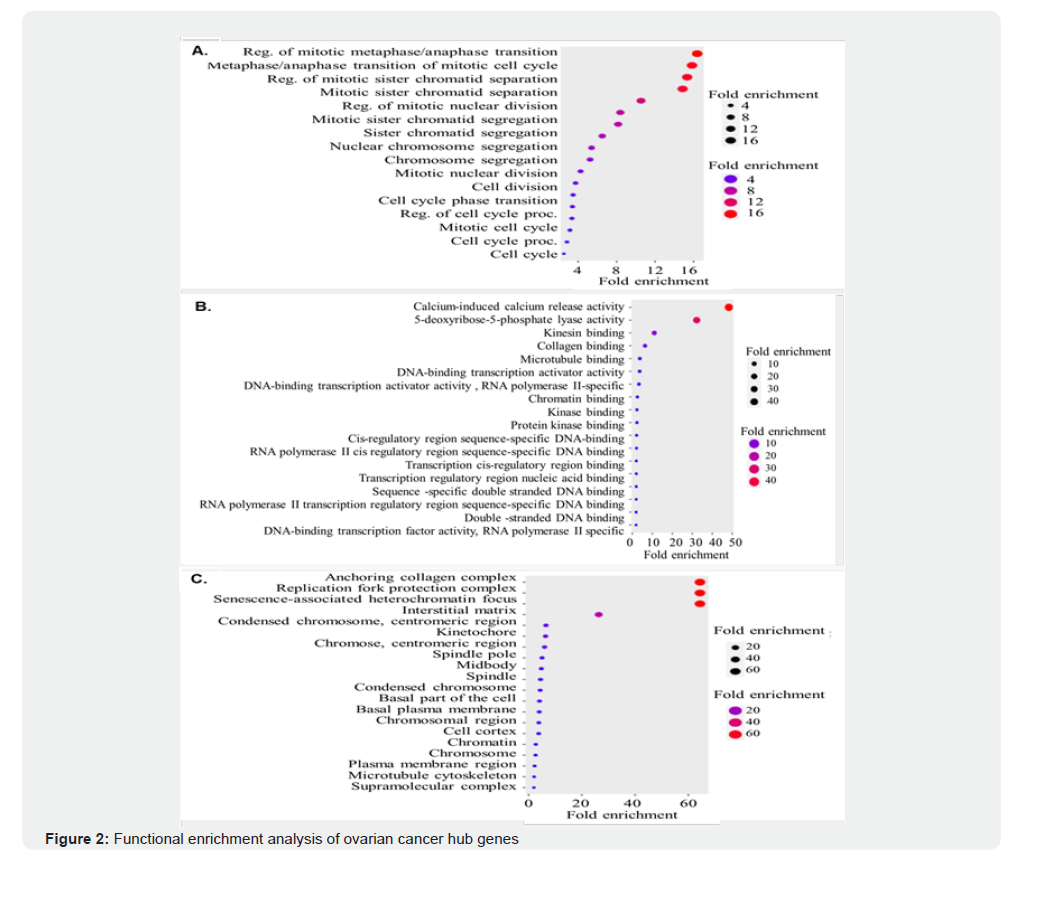
In the MF category, overlapping DEGs showed marked enrichment in activities related to calcium-induced calcium release and 5-deoxy-5-ribose-phosphate lyase, with fold enrichment exceeding 4 (Figure 2B). For the cellular component, DEGs were significantly associated with the anchoring collagen complex, replication fork protection complex, and senescence-associated heterochromatin foci (Figure 2C), suggesting a potential role in maintaining genomic integrity and cellular senescence. KEGG pathway enrichment analysis demonstrated that these DEGs were primarily involved in pathways such as tyrosine, glycineserine- threonine, retinol, cell cycle, and drug metabolism. These findings highlight key molecular pathways that may contribute to OC progression and offer potential therapeutic targets for intervention.
Gene Ontology (GO) enrichment analysis was performed to characterize the biological relevance of the identified hub genes. (A) Biological Process (BP) terms were significantly enriched in pathways regulating cell cycle progression, mitotic nuclear division, and chromosome segregation, highlighting strong involvement in mitotic control. (B) Molecular Function (MF) enrichment revealed associations with DNA-binding transcriptional activity, protein kinase binding, chromatin binding, and regulatory sequence-specific DNA binding, underscoring their role in transcriptional regulation and cell-cycle checkpoint control. (C) Cellular Component (CC) analysis indicated strong enrichment in chromosomal regions, spindle apparatus, kinetochore, replication fork protection complexes, and anchoring collagen complexes, reflecting their localization to structures critical for mitosis, chromosomal stability, and cell division. Node size corresponds to fold enrichment, with color intensity reflecting degree of significance. The PPI network was constructed using differentially expressed genes (DEGs) significantly associated with ovarian cancer, comprising 251 nodes and 1,019 edges with an average node degree of 8.12 and clustering coefficient of 0.436 (expected edges: 271; PPI enrichment p < 0.001). FOXM1 (highlighted in yellow) emerges as a central hub interacting with key regulators of the cell cycle, mitosis, and transcriptional control, including CDK1, CCNB1, CCNB2, AURKB, CENPA, CENPF, MYC, and EP300. Red edges denote stronger confidence interactions, while blue edges represent moderate interactions. The high connectivity of FOXM1 underscores its role as a master regulator of oncogenic signaling and validates its potential as a therapeutic target in ovarian cancer. The PPI network analysis identified 251 nodes and 1,019 edges, underscoring the complexity of the gene interactions involved in OC. Among these, the module genes AURKB, CCNB1, CCNB2, CDK1, CENPA, CENPF, FOXM1, and NEK2 emerged as the top eight hub genes, exhibiting the highest node degree (Supplementary Figure 1) and, thus playing a central role in network functionality. These hub genes were highlighted because of their significant involvement in key biological processes, particularly those related to cell cycle regulation and mitotic control.
Furthermore, the most crucial PPI network module was identified using the MCODE algorithm, which identified highly interconnected gene clusters. Pathway enrichment analysis of this top module revealed that these genes were predominantly enriched in critical pathways such as the p53 signaling pathway, progesterone-mediated oocyte maturation, oocyte meiosis, cell cycle, and cellular senescence (Table 2). These findings suggest that these pathways may serve as pivotal regulators of OC progression and offer potential therapeutic avenues for targeted intervention.
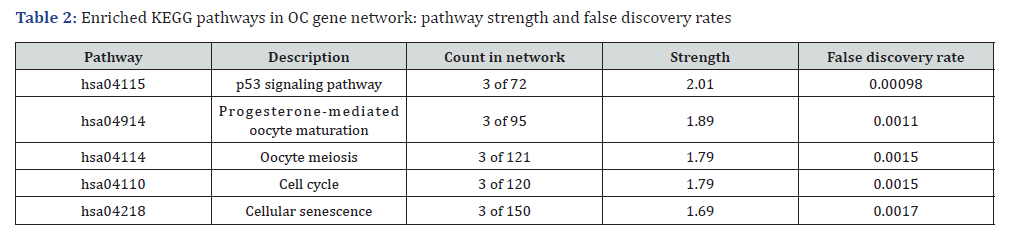
Footnote: The table lists the enriched KEGG pathways identified in the OC gene network. The “Count in network” column represents the number of genes involved in each pathway out of the total network size. “Strength” refers to the degree of enrichment, while the “False Discovery Rate” indicates the likelihood of false positives, with lower values representing higher confidence in the enrichment.
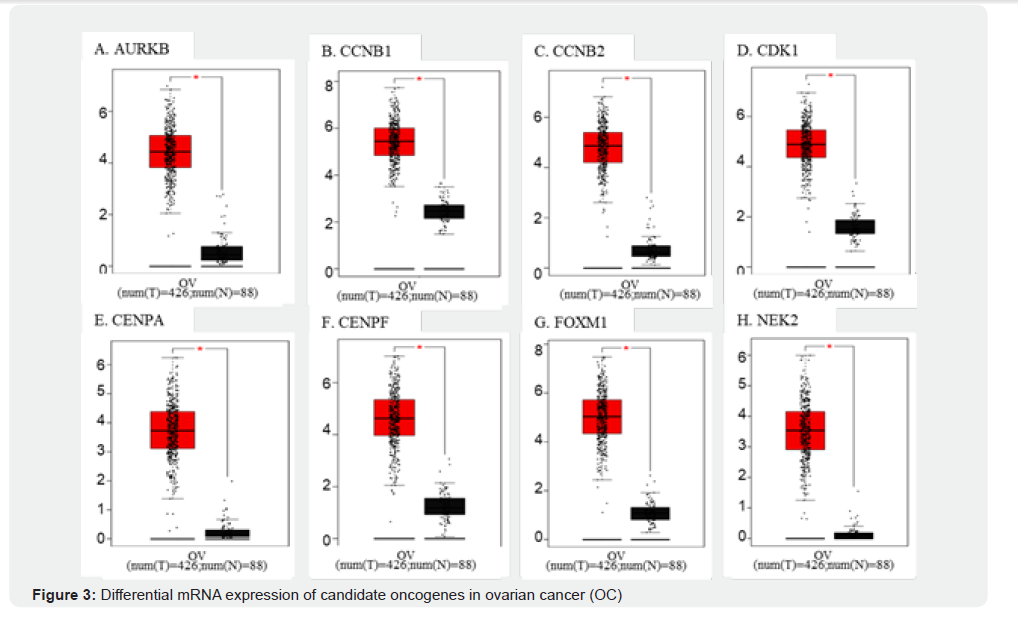
Using GEPIA and Oncomine, we confirmed that AURKB, CCNB1, CCNB2, CDK1, CENPA, CENPF, FOXM1, and NEK2 were significantly overexpressed in OC tissues compared to normal tissues (Figure 3A-H). Overall survival (OS) curves for ovarian cancer patients were generated according to high (red) and low (black) mRNA expression levels of (A) AURKB, (B) CCNB1, (C) CCNB2, (D) CDK1, (E) CENPA, (F) CENPF, (G) FOXM1, and (H) NEK2. Hazard ratios (HR) with 95% confidence intervals and logrank p-values are shown. High expression of CCNB1, CCNB2, CDK1, CENPA, CENPF, FOXM1, and NEK2 was significantly associated with worse OS, underscoring their prognostic value in ovarian cancer. Elevated levels of CCNB1, TOP2A, NUSAP1, NCAPG, KIF20A, and DLGAP5 were also linked to worse OS in patients (Supplementary Figure 2A-H), emphasizing their clinical relevance. The Human Protein Atlas further validated the high protein expression levels of these genes in OC. Notably; FOXM1 has emerged as a key oncogene with strong overexpression, prognostic significance, and drug-targeting potential, making it a prime candidate for therapeutic development in OC. As shown in (Table 3), AURKB was predominantly localized in the nuclear region, with a high expression rate of 7/12 (58.3%) in serous ovarian cancer (SOC) samples, which was significantly higher than that in benign serous ovarian cystadenoma (5/12, 41.6%). CCNB1 and CCNB2 were primarily located in the cytoplasmic/membranous region, with low expression rates of 9/12 (58.3%) and 5/12 (58.3%) in SOC, respectively, both of which were higher than those in the benign samples. CDK1 showed nuclear and cytoplasmic localization, with a low expression rate of 4/11 (58.3%) in SOC compared to 5/12 (41.6%) in benign cystadenomas. CENPA was localized to the nucleus, with high expression in 6/12 (58.3%) SOC samples, which was higher than that in benign samples. CENPF exhibited both nuclear and cytoplasmic localization, with a low expression rate of 8/12 (58.3%) in SOC. Notably, FOXM1 was highly expressed in 10/11 (90.9%) SOC samples, which was significantly higher than that in benign cystadenoma (1/11, 9.1%, P<0.05), with localization in both the nucleus and the cytoplasmic membrane.
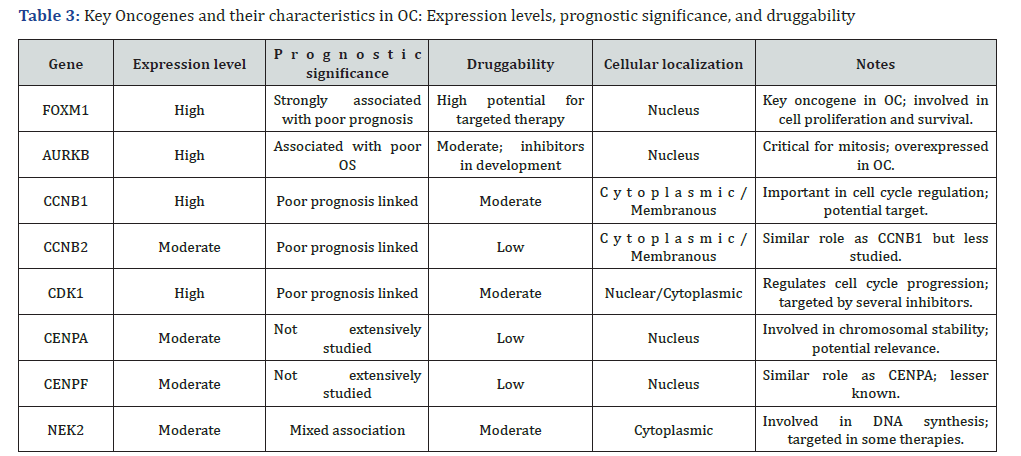
Footnote: The table summarizes critical genes involved in OC, detailing their expression levels, prognostic significance, druggability potential, and cellular localization. FOXM1, AURKB, and CCNB1 are highlighted for their strong association with poor prognosis and potential as therapeutic targets. Druggability refers to the feasibility of targeting these genes with existing or developmental therapeutic compounds.
Box plots show the relative expression levels of (A) AURKB, (B) CCNB1, (C) CCNB2, (D) CDK1, (E) CENPA, (F) CENPF, (G) FOXM1, and (H) NEK2 in ovarian cancer tissues (red, n = 426) compared with normal ovarian tissues (black, n = 88), based on GEPIA and Oncomine analyses. All genes were significantly overexpressed in OC tissues (p < 0.05, indicated by red asterisks). These findings highlight the aberrant activation of cell-cycle– related regulators and underscore their potential roles in ovarian cancer pathogenesis. In OC single-cell functional state analysis, FOXM1 was identified as the most critical gene among the eight genes analyzed, demonstrating the strongest correlation with cell cycle regulation (score: 0.83) and proliferation (score: 0.79) (Figure 4A-4H). These associations indicate that FOXM1 plays a central role in driving OC tumor progression by promoting rapid cell division and facilitating cell cycle transition. Its significant involvement in these essential cancer processes underscores FOXM1 as a top priority for therapeutic targeting, particularly in treatments aimed at disrupting uncontrolled tumor proliferation. Other genes, such as AURKB, CCNB1, CCNB2, CDK1, CENPA, CENPF, and NEK2 also contribute to functional states linked to the cell cycle and proliferation, but their impact is less pronounced. For instance, CDK1 and AURKB are involved in cell cycle regulation, whereas CCNB1 and CCNB2 are involved in cell division and mitosis. Despite these roles, FOXM1’s dominant influence on both the cell cycle and proliferative activity in OC positions it as the most important gene in this cohort, making it a primary target for potential therapeutic interventions.
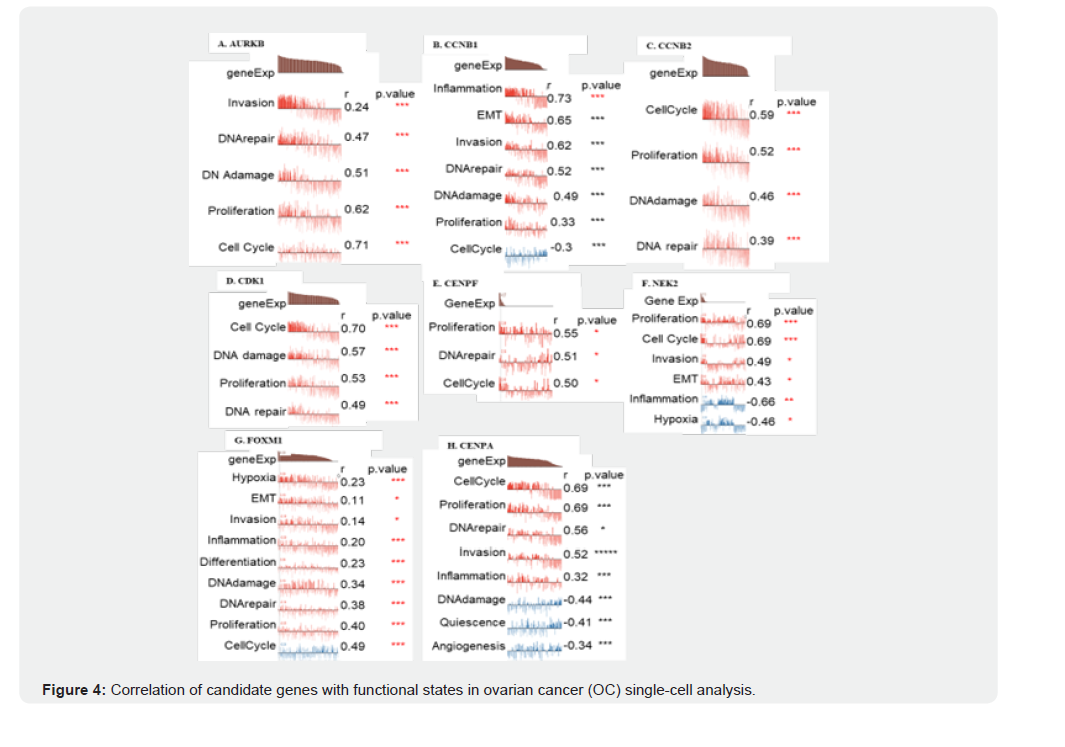
Single-cell transcriptomic functional state analysis was performed to evaluate the association of eight genes (AURKB, CCNB1, CCNB2, CDK1, CENPF, NEK2, FOXM1, and CENPA) with key cancer-related processes. (A–H) Correlation plots depict the relationship between gene expression levels and functional states, including cell cycle, proliferation, DNA damage, DNA repair, invasion, epithelial–mesenchymal transition (EMT), inflammation, angiogenesis, hypoxia, and quiescence. Among the analyzed genes, FOXM1 demonstrated the strongest correlation with cell cycle regulation (r = 0.83) and proliferation (r = 0.79), underscoring its central role in driving tumor progression by promoting rapid cell division and facilitating cell cycle transitions. Although other genes, including AURKB, CDK1, CCNB1, CCNB2, CENPF, NEK2, and CENPA, also exhibited positive associations with cell cycle and proliferative states, their impact was less pronounced compared to FOXM1. These findings highlight FOXM1 as a dominant regulator of proliferative activity in OC, making it a promising therapeutic target for interventions aimed at disrupting uncontrolled tumor growth.
The table highlights FOXM1 as the best candidate among the eight genes based on various criteria such as expression level, prognostic significance, druggability, and localization. This table includes other genes for comparison, emphasizing the advantages of FOXM1.
FOXM1 is considered for further analysis because of its strong association with poor prognosis in OC and its high druggability potential. As a key oncogene, FOXM1 plays a crucial role in promoting cell proliferation and survival and is a critical factor in cancer progression. Its high expression levels and nuclear localization highlight its relevance in tumor biology. Given its involvement in these fundamental cellular processes and its potential as a therapeutic target, FOXM1 represents a promising candidate for developing targeted treatments for OC, warranting further investigation. This figure illustrates the differential expression of FOXM1 between tumor and normal tissues and further evaluates its potential diagnostic utility through specificity and sensitivity analyses based on normal tissue expression thresholds. Analysis of FOXM1 expression revealed a significant elevation in OC tissues compared to normal controls. The boxplot (left panel) demonstrated markedly higher FOXM1 transcript levels in tumors, with a clear upward shift in expression distribution and a borderline significant difference (P = 5.91 × 10⁻²). To further assess the discriminatory power of FOXM1, tumor expression levels were compared against different percentile-based cutoffs derived from normal tissues (right panel). Across minimum, first quartile (Q1), median, and third quartile (Q3) thresholds of normal FOXM1 expression, 100% of tumor samples exhibited higher expression, indicating robust overexpression across the OC cohort. Even at the maximum cutoff of normal expression, 40% of tumors remained above the threshold, emphasizing that a substantial subset of OC harbors FOXM1 upregulation beyond physiological variation. (A) Boxplot showing FOXM1 expression significantly elevated in tumor tissues compared with normal controls (P = 5.91 × 10⁻²). (B) Proportion of tumors exceeding FOXM1 expression cutoffs derived from normal tissues. Across minimum, Q1, median, and Q3 thresholds, 100% of tumors showed higher expression, while 40% remained elevated even above the maximum normal cutoff. Specificity increased progressively with higher cutoffs, reaching 1.0 at the maximum threshold. Specificity analysis showed a progressive increase from ~0.55 at the minimum cutoff to ~1.0 at the maximum cutoff, confirming FOXM1’s strong discriminatory potential between tumor and normal states (Supplementary Figure 3). These findings collectively highlight FOXM1 as a consistently elevated gene in OC, supporting its role as a tumorassociated driver and a candidate diagnostic biomarker.
In summary, this figure underscores FOXM1’s potential as a tumor biomarker, with consistent overexpression across cancer samples. Its diagnostic value depends on the cut-off strategy; lower thresholds offer high sensitivity but lower specificity, whereas higher thresholds increase specificity at the expense of sensitivity. These findings suggest that FOXM1, either alone or in combination with other markers, may be a useful candidate for cancer detection and stratification. FOXM1 (Forkhead box protein M1) is a critical transcription factor that regulates cell cycle genes essential for DNA replication and mitosis, making it a key player in OC progression. The PPI network surrounding FOXM1 highlights its central role in regulating critical mitotic processes, as evidenced by its association with key proteins such as CDK1 and PLK1 (Supplementary Figure 1). CDK1 is pivotal for G2 to M phase transition, whereas PLK1 facilitates various functions during mitosis, including spindle assembly and mitotic exit. Additionally, FOXM1 interacts with cyclin CCNB1 and CCNA2, which control the transition to the G2/M and G1/S phases. These interactions emphasize FOXM1’s function in orchestrating cell cycle checkpoints and maintaining proliferative signaling in OC cells.
Moreover, FOXM1 interactions extend beyond traditional cell cycle regulators to include proteins such as CTNNB1 and EP300, suggesting potential crosstalk with signaling pathways such as Wnt and epigenetic regulation of gene expression. The connection between FOXM1 and BIRC5 (survivin) indicated its role in promoting cancer cell survival by inhibiting apoptosis, further confirming its importance in OC. Given these interactions and FOXM1’s overarching influence on cell proliferation and survival, targeting FOXM1 and its network partners could represent a promising therapeutic strategy for disrupting the aggressive proliferation of OC cells, positioning FOXM1 as a vital candidate for further research and clinical intervention. GO and KEGG pathway analyses of the FOXM1 PPI network in OC revealed significant enrichment in several BP, cellular components, and molecular functions relevant to cancer biology. In the BP category, FOXM1-associated genes were particularly enriched in pathways related to the G2/M transition of the mitotic cell cycle (p = 1. x 10- 10), cell cycle G2/M phase transition (p-value: 1.6 x 10-10), and mitotic cell cycle checkpoint signalling (p-value: 5.9 x 10-7). These findings indicated that FOXM1 is involved in cell cycle progression and checkpoint regulation, which are critical mechanisms in cancer proliferation and tumorigenesis (Figure 5A).
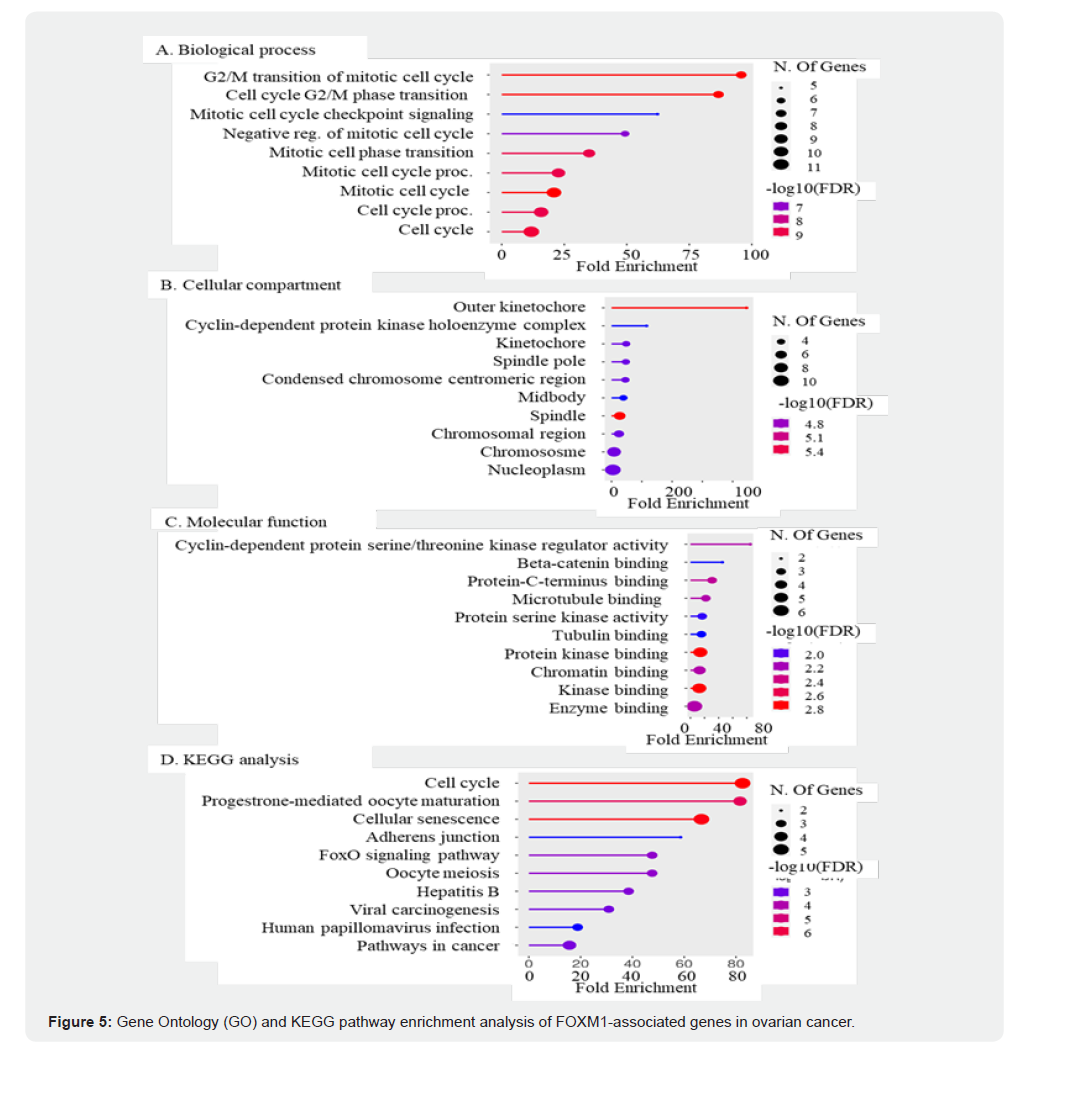
In the Cellular Component (CC) category, the genes associated with FOXM1 were significantly enriched in the outer kenetockore (p = 2.2 x 10-6), cell cycle-dependent protein kinase holoenzyme complex (p = 3.0 x 10-5), and nucleoplasm (p = 2. x 10-5). Enrichment in the nucleoplasm (p = 2.3 x 10-5) further underscores their role in altered nuclear dynamics and transcriptional regulation, a hallmark of cancer progression and metastasis (Figure 5B). This association indicates that FOXM1 and its interacting partners not only play essential roles in cell cycle control but also contribute to the regulatory networks governing gene expression within the nucleus.
(A) Biological Process (BP) analysis revealed significant enrichment in cell cycle–related pathways, including G2/M transition of the mitotic cell cycle, mitotic checkpoint signaling, and regulation of mitotic processes. (B) Cellular Component (CC) analysis showed enrichment in outer kinetochore, cyclindependent protein kinase holoenzyme complex, spindle pole, and nucleoplasm, underscoring FOXM1’s nuclear and mitotic functions.
(C) Molecular Function (MF) analysis indicated significant enrichment in cyclin-dependent protein serine/threonine kinase regulator activity, β-catenin binding, and protein C-terminus binding, highlighting FOXM1’s role in cell cycle regulation and transcriptional control. (D) KEGG pathway analysis demonstrated enrichment in cell cycle, progesterone-mediated oocyte maturation, cellular senescence, adherens junction, and pathways in cancer, emphasizing FOXM1’s central involvement in ovarian tumorigenesis and progression. Dot size indicates the number of enriched genes, while color represents significance level (–log10 FDR).
In the MF category, the analysis revealed significant enrichment in cyclin-dependent protein serine/threonine kinase regulator activity (p-value: 5.7 x 10-3), beta-catenin binding (1.2 x 10-2), and Protein C-terminus binding (p-value: 4.8 x 10-3). This emphasizes the role of these MFs in regulating critical biological processes that maintain cellular homeostasis and tissue integrity, which are frequently targeted in cancer therapies to disrupt tumor growth and survival (Figure 5C). FOXM1 serves as a critical regulator of MFs essential for cell cycle control and signaling, influencing key pathways that maintain cellular homeostasis and tissue integrity, thereby positioning it as a promising therapeutic target for OC. KEGG pathway analysis reinforced these findings, revealing substantial enrichment of several pathways that are crucial for cancer progression. Cell cycle (p-value: 1.3 x 10-7), progesterone-mediated oocyte maturation (2.5 x 10-6), cellular senescence (p-value: 1.9 x 10-7), and adherens junction (p-value: 3.2 x 10-3) (Figure 5D). These findings indicate the involvement of the gene set in critical processes such as cell proliferation, hormonal regulation of ovarian functions, cellular aging, and maintenance of cell adhesion within the tumor microenvironment, highlighting their collective contribution to tumorigenesis and cancer progression. Additionally, enrichment in the FoxO signaling pathway (p-value: 3.7 x 10-4) highlights the role of this gene set in regulating oxidative stress responses and apoptosis, both of which are essential for regulating cancer cell survival and proliferation. This suggested that FOXM1 and its associated genes may influence the balance between cell death and survival, thereby playing a critical role in OC progression and therapeutic resistance. Together, GO and KEGG pathway analyses of the FOXM1 PPI network provided a comprehensive understanding of the functional roles of FOXM1-associated genes in OC, revealing key biological processes and signaling pathways that could be exploited for therapeutic interventions, particularly through targeted and combination therapy approaches. These insights emphasize the potential of FOXM1 as a strategic target for the development of novel therapeutic strategies aimed at disrupting molecular mechanisms underlying OC progression.
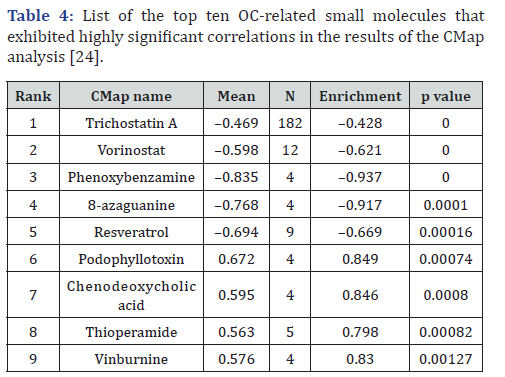
According to the p-value analysis, the top eight small molecules correlated with OC gene expression changes are listed in Table 5. Six of these molecules exhibited a negative correlation, indicating their potential tumor-inhibitory effects in clinical settings. These four molecules were positively correlated. Among those with significant negative correlations (P<0.05), trichostatin A, vorinostat, 8-azaguanine and phenoxybenzamine suggested that these compounds may suppress tumorigenic pathways. CMap analysis identified 19 small molecules significantly associated with OC (Table 4). Trichostatin A and vorinostat have emerged as the most promising therapeutic inhibitors, with highly negative enrichment scores and extremely low P-values, indicating a strong potential for clinical use. Phenoxybenzamine and 8-azaguanine also showed significant negative correlations, further supporting their relevance to the inhibition of OC-related pathways. In contrast, podophyllotoxin, chenodeoxycholic acid, and thioperamide exhibited positive enrichment scores, implying their potential roles in activating or supporting key OC biological processes. These findings suggest that these molecules hold promise for therapeutic development and as investigative tools for OC research.
The table highlights the significant overexpression of key genes (FOXM1, CCNB1, AURKB, CDK1, CENPA, CCNB2, CENPF, and NEK2) in malignant OC compared with that in benign and borderline tumors. FOXM1, with a highly significant p-value (< 0.00001), was overexpressed in 77.1% of malignant cases, underscoring its critical role in tumor proliferation and aggressiveness. Similarly, genes such as CCNB1, AURKB, and CDK1, with p-values of 0.00036, 0.00045, and 0.00069, respectively, were strongly involved in cell cycle regulation, particularly in the G2/M transition and mitotic processes. Differential expression of these genes across malignancy levels not only marks them as valuable diagnostic biomarkers but also highlights their potential in prognostic assessments by correlating their expression with disease progression and patient outcomes. The therapeutic potential of these genes is attributed to their direct involvement in vital cellular processes. FOXM1, AURKB, and CDK1 are particularly promising druggable targets, where inhibitors aimed at halting cell cycle progression or disrupting mitotic function could effectively impair tumor growth. Given their critical roles in tumor biology, targeting these genes offers opportunities for the development of precise mechanism-based treatments for OC. Their high prognostic value combined with their draggability positions them as key targets for both personalized medical approaches and novel therapeutic interventions, ultimately enhancing the efficacy of OC treatment strategies. FOXM1, CCNB1, and AURKB have emerged as top performers in malignancy, with FOXM1 showing the highest significance and potential as a therapeutic target because of its strong association with malignancy. CCNB1 and AURKB are also significantly overexpressed in malignant tumors, suggesting their role in tumor progression and making them promising targets for targeted therapies. CDK1 and CENPA are notably expressed in malignant cases, and may be valuable for both diagnostic and therapeutic applications. CCNB2 and CENPF offer slightly less pronounced results, but still represent potential biomarkers and therapeutic targets. NEK2, although statistically significant, requires further validation to incomplete data (Table 5).
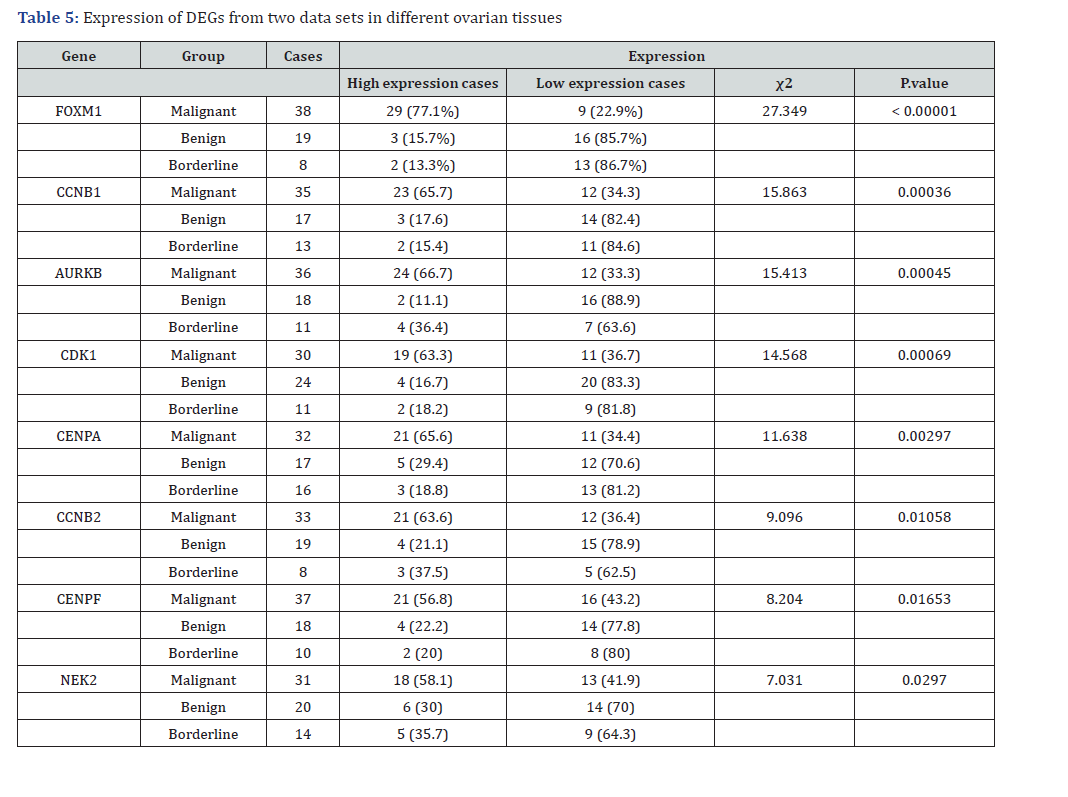
Footnote: The table shows gene expression differences (high vs. low) across malignant, benign, and borderline tumor groups. χ² tests compare expression distributions, with significant P-values (< 0.05) indicating association between gene expression and tumor type. Percentages reflect proportions within each group. Missing values (e.g., NEK2 benign/borderline) suggest data formatting gaps.
The identification of the binding cavity in which FOXM1 interacts with small molecules is pivotal for the discovery of novel FOXM1 inhibitors via structure-based virtual screening. The crystal structure of FOXM1 (PDB ID: 3G73) retrieved from the RCSB protein database was used to generate the receptor model for molecular docking. Previous studies have mapped the binding site of FOXM1 where small molecules are predicted to interact, as shown in (Figure 6A). The surface-binding mode of FOXM1 is depicted in (Figure 6B), highlighting its interaction dynamics. (A) 3D surface representation of FOXM1 binding pocket showing the ligand docked at the active site. The binding surface is colored according to electrostatic potential, with the ligand shown in stick representation. Key binding interactions, including hydrogen bonds, π–π stacking, and hydrophobic contacts, are highlighted. (B) 2D interaction diagram of the ligand–FOXM1 complex depicting the specific amino acid residues involved in ligand binding. Conventional hydrogen bonds (green), π–π interactions (purple), alkyl interactions (orange), and van der Waals forces (grey) are indicated. Notable interacting residues include ARG236, LYS278, ALA278, PRO279, GLY280, TRP281, and ASN283, which stabilize ligand binding within the FOXM1 active site. Key residues, particularly ARG236 and TYR272, were identified as critical for the binding affinity between FOXM1 and small molecules, as shown by the docking pose (Supplementary Figures 4A and 4B). These amino acids contribute to the structural integrity and functional capacity of the binding site, which was used for the subsequent virtual screening of potential inhibitors. Our study commenced with a structure-based virtual docking screen of the NCI Diversity Set 2, which encompasses 265,242 chemically diverse compounds (Figure 6C). Using AutoDock Vina (ADV), we docked 1,880 of these compounds to FOXM1. The compounds were ranked based on their predicted binding free energies (ΔG_ADV), which varied from −2.8 to −7.8 kcal/mol. Notably, 13.7% of the compounds exhibited binding energies lower than −6.0 kcal/mol, surpassing the benchmark set by FDI-6 (Figure 6D).
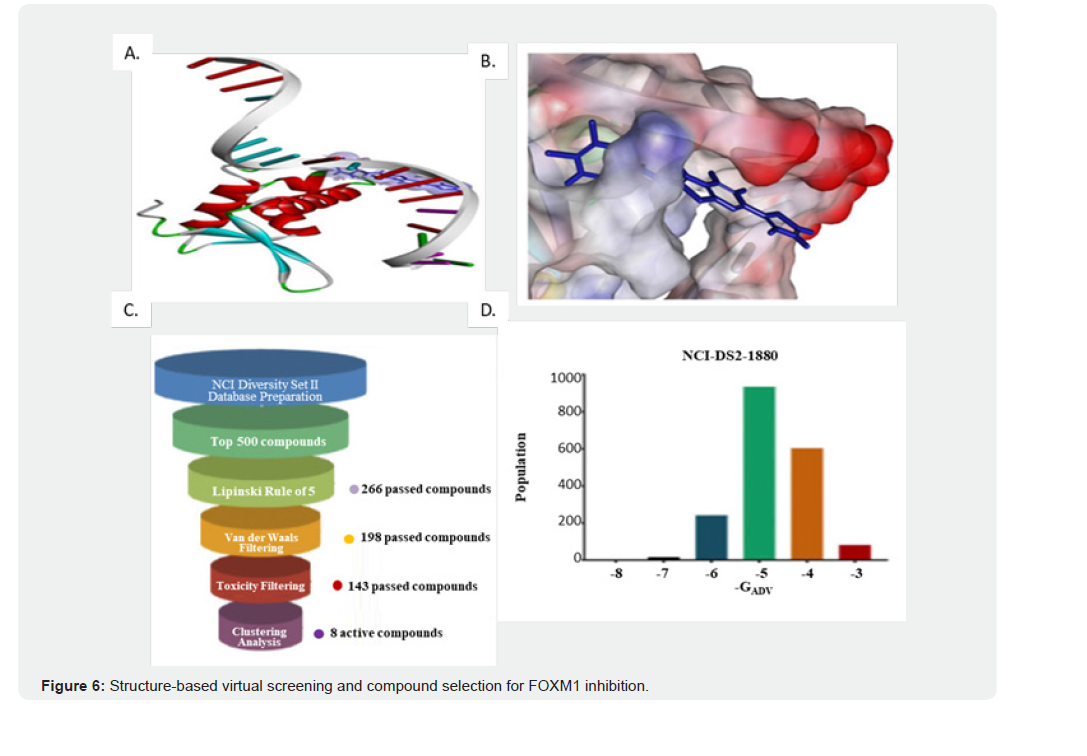
Ribbon representation of FOXM1 protein structure showing its DNA-binding domain. (B) Molecular docking visualization depicting the binding pose of a screened compound within the FOXM1 active site. (C) Workflow of virtual screening and filtering pipeline: from NCI Diversity Set II, the top 500 compounds were selected, followed by successive filtering steps including Lipinski’s Rule of Five (266 compounds), van der Waals filtering (198 compounds), toxicity filtering (143 compounds), and clustering analysis, yielding 8 candidate active compounds. (D) Distribution of docking free energy scores (ΔGADV) across screened compounds, highlighting the strong binding profile of NCI-DS2-1880. To refine the selection process and discard unsuitable candidates, a threestep filtering process was applied. First, we narrowed down to the top 500 compounds based on their binding energies and then assessed them using Lipinski’s “Rule of Five” and the Veber rule to ensure drug-like properties. Finally, we utilized the “toxicity prediction” tool in Discovery Studio (DS) to exclude compounds with potential carcinogenic, mutagenic, or teratogenic effects. This rigorous filtering resulted in a final set of 25 drug-like compounds suitable for further evaluation.
Subsequently, we conducted a comprehensive evaluation of the binding energy data, key residue interactions at binding sites, and structural diversity. From this refined library, we selected seven compounds for future in vitro biological evaluations using their DS virtual screening scores. Docking analysis identified the compound ZINC13597767 (TV4) as the most promising candidate, demonstrating the most favorable interaction with the FOXM1 target. This compound achieved the lowest CDOCKER energy of −18.1145 kcal/mol, forming stable hydrogen bonds with ARG236 and TYR272, which were crucial for its strong binding affinity. Conversely, compound ZINC00001087 (TV5) displayed a highly unfavorable interaction, characterized by a positive CDOCKER energy of 30.7268 kcal/mol, indicating poor binding and potential steric clashes. Several other compounds, such as ZINC00039221 (TV3) and ZINC00031410 (TV6), also exhibited negative CDOCKER energies, and successfully formed hydrogen bonds with ARG236 and TYR272. These results underscore the critical role of these amino acid residues in stabilizing ligand binding and highlight their significance in the design of effective FOXM1 inhibitors (Table 6).
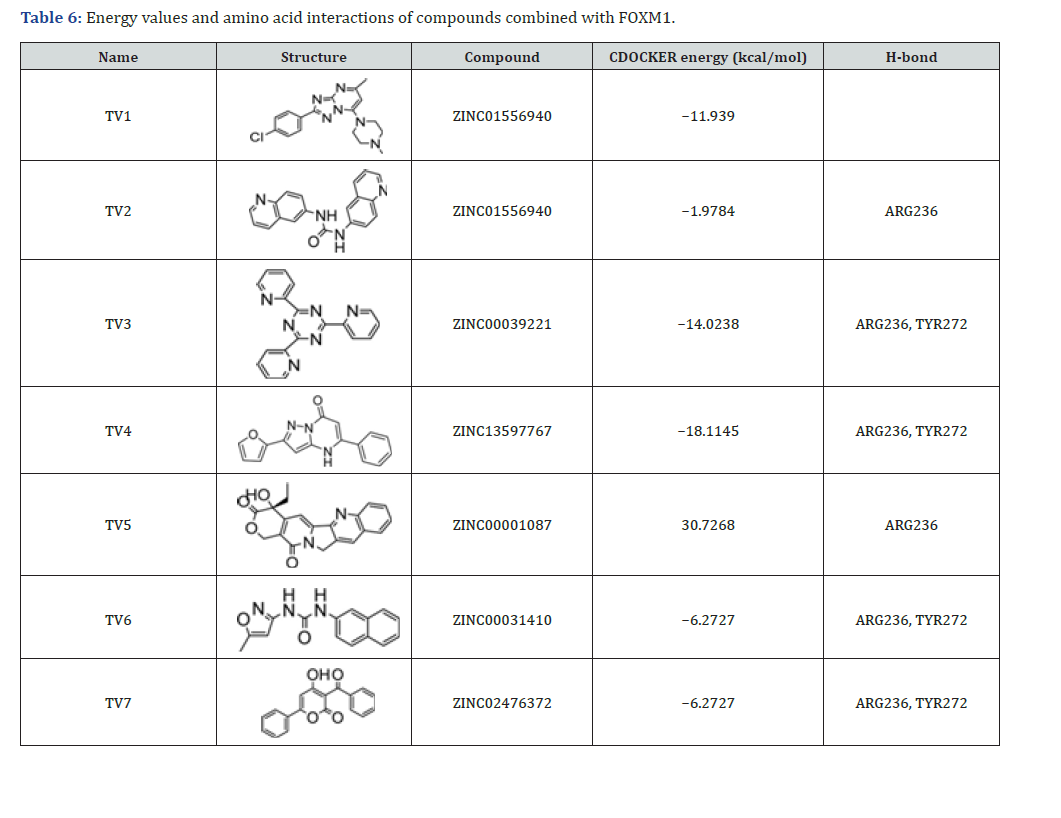
Footnote: Negative CDOCKER energy values indicate favorable binding. H-bond interactions with key residues (ARG236, TYR272) are shown. TV5’s positive energy suggests unstable binding.
The results from the CMap and docking analyses collectively enhance the evaluation of small molecules for OC therapy. CMap analysis identified Trichostatin A and Vorinostat as promising candidates owing to their significant negative correlation with OC, indicating their potential as effective therapeutic inhibitors. Concurrently, docking analysis provided insights into the binding affinities of specific compounds to FOXM1, with ZINC13597767 emerging as the most favorable candidate, as evidenced by its low CDOCKER energy and the formation of hydrogen bonds with crucial residues ARG236 and TYR272. This synergy between CMap’s disease correlation insights and the docking analysis’s binding potential assessments refines the selection of compounds, validating their suitability for further in vitro evaluation, and supporting their advancement in therapeutic development.
Discussion
Summary of Main Results
The primary objective of this study was to identify key driver genes in OC pathogenesis through a multi-omics approach. Our main finding is the identification of 4,204 DEGs, from which 16 were significantly associated with poor overall survival. Among these, the transcription factor FOXM1 emerged as the most critical oncogenic driver, demonstrating strong nuclear expression in 90.9% of serous carcinomas compared to 9.1% in benign tissues. Secondary analyses confirmed the role of this gene set in cell cycle processes and identified ZINC13597767 as a promising FOXM1 inhibitor through virtual screening.
Results in the Context of Published Literature
Our findings strongly align with and extend the work of TCGA network, which established the landscape of molecular alterations in high-grade serous OC [27]. The central role of FOXM1 we identified is consistent across diverse datasets [53] and is mechanistically supported by its known activation by mutant p53, a nearly universal event in this cancer subtype [28]. The overexpression of our identified hub genes (FOXM1, AURKB, CCNB1, CDK1) and their association with aggressiveness and chemoresistance have been individually reported [29]. However, our study integrates these findings into a cohesive network, positioning FOXM1 as a master regulator. While previous studies have suggested FOXM1’s diagnostic utility, our ROC analysis provides a quantitative assessment of its discriminatory power [30]. Furthermore, our virtual screening result for ZINC13597767 is novel, though it is supported by the known efficacy of other HDAC inhibitors like vorinostat in suppressing FOXM1 activity [31].
Strengths and Weaknesses
A key strength of this study is the integrative multi-omics methodology, which combined DEG analysis, prognostic validation, functional enrichment, PPI network construction, and experimental wet-lab validation to robustly identify and prioritize FOXM1. This approach mitigates the risk of false discoveries common in single-platform analyses. The use of virtual screening to identify a novel putative FOXM1 inhibitor (ZINC13597767) is strength, providing a direct translational pathway for future research. The main weakness is the lack of comprehensive in vivo functional validation for FOXM1 and the top candidate compound. Furthermore, the study primarily focused on the highgrade serous subtype, and the findings may not be generalizable to other OC histotypes (e.g., clear cell, mucinous), which have distinct molecular profiles and clinical behaviors [32]. The diagnostic specificity of FOXM1 alone remains variable, and the pharmacokinetics and potential toxicity of the identified inhibitor candidates remain to be fully characterized [33].
Implications for Practice and Future Research
The most impactful contribution of this study is the systematic prioritization of FOXM1 as a high-value master regulator and therapeutic target in OC, supported by a novel computational drug candidate. For clinical practice, measuring FOXM1 expression could enhance risk stratification and its integration with CA-125 may improve early detection strategies [34]. For future research, the immediate priority is the in vitro and in vivo validation of ZINC13597767 to confirm its efficacy and safety as a FOXM1 inhibitor. Subsequently, exploring its synergy with existing agents (e.g., PARP inhibitors, platinum chemotherapy) and mitotic kinase inhibitors (e.g., barasertib, dinaciclib) is warranted. Future work must also expand to non-serous subtypes to determine the pan- OC applicability of this target. Finally, given emerging evidence that FOXM1 modulates the tumor immune microenvironment, investigating its inhibition in combination with immunotherapy presents a compelling and novel research direction.
Conclusion
In conclusion, our integrated analysis highlights FOXM1 as a central driver of OC pathogenesis, with profound diagnostic, prognostic, and therapeutic implications. Dysregulation of the cell cycle and p53 signaling pathways, coupled with the identification of novel small-molecule inhibitors, provides a strong foundation for targeted drug development. By bridging bioinformatics insights with translational research, this study offers a roadmap for improving OC outcomes, a disease that urgently requires innovative therapeutic strategies. This conclusion aligns with the growing body of research that recognizes FOXM1 as a therapeutic vulnerability across multiple tumor types, including OC. The convergence of pathway analysis, experimental validation, and drug screening echoes the strategy proposed in precision oncology frameworks [35] suggesting that such integrative approaches may accelerate clinical translation in high-burden cancers such as OC.
Author Contributions
T.W., M.B. and M. S. contributed to the conceptualization, data curation, and formal analysis of the article. T.W. was responsible for writing the original draft, preparation, visualization, and methodology. M.B. reviewed and edited the manuscript. M.B. provided supervision, project administration, and resources, and secured funding for the study. All the authors have read and approved the final version of the manuscript.
Funding
This work was supported by the Wolkite University research and community service Vice president office.
Acknowledgments
We acknowledge the Wolkite University, Ethiopia and Institute of Biopharmaceutical and Health Engineering, Tsinghua Shenzhen International Graduate School, China for administrative support.
References
- Hong MK, DC Ding (2025) Early Diagnosis of Ovarian Cancer: A Comprehensive Review of the Advances, Challenges, and Future Directions. Diagnostics (Basel) 15(4): 406.
- Marth C, Abreu MH, Andersen KK, Aro KM, Batarda ML, et al., (2022) Real-life data on treatment and outcomes in advanced ovarian cancer: An observational, multinational cohort study (RESPONSE trial). Cancer 128(16): 3080-3089.
- Chacko N, R Ankri (2024) Non-invasive early-stage cancer detection: current methods and future perspectives. Clinical and Experimental Medicine 25(1): 17.
- Pawłowska A, Rekowska A, Kuryło W, Pańczyszyn A, Kotarski J, et al., (2023) Current Understanding on Why Ovarian Cancer Is Resistant to Immune Checkpoint Inhibitors. Int J Mol Sci 24(13).
- Satam H, Joshi K, Mangrolia U, Waghoo S, Zaidi G, et al., Next-Generation Sequencing Technology: Current Trends and Advancements. Biology (Basel) 12(7): 997.
- Tzec-Interián JA, D González-Padilla, EB Góngora-Castillo (2025) Bioinformatics perspectives on transcriptomics: A comprehensive review of bulk and single-cell RNA sequencing analyses. Quantitative Biology 13(2): e78.
- Zhou X, Chen Y, Zhang Z, Miao J, Chen G, et al., (2024) Identification of differentially expressed genes, signaling pathways and immune infiltration in postmenopausal osteoporosis by integrated bioinformatics analysis. Heliyon, 2024. 10(1): e23794.
- Wolde T, Bhardwaj V, Md Reyad-Ul-Ferdous, Qin P, Pandey V (2024) The Integrated Bioinformatic Approach Reveals the Prognostic Significance of LRP1 Expression in Ovarian Cancer. International Journal of Molecular Sciences 25(14): 7996.
- Liao L, Han W, Shen Y, Shen G (2024) Comprehensive analysis of aberrantly methylated differentially expressed genes and validation of CDC6 in melanoma. Journal of Cancer Research and Clinical Oncology 150(7): 362.
- Rosati D, Palmieri M, Brunelli G, Morrione A, Iannelli F, et al., (2024) Differential gene expression analysis pipelines and bioinformatic tools for the identification of specific biomarkers: A review. Comput Struct Biotechnol J, 23: 1154-1168.
- Wang Z, Zhang B, Li D, Qi X, Chijin Zhang, et al., (2021) Bioinformatic analysis of key pathways and genes involved in pediatric atopic dermatitis. Bioscience Reports 41(1).
- Majeed A, Mukhtar S, (2023) Protein-Protein Interaction Network Exploration Using Cytoscape. Methods Mol Biol. 2690: 419-427.
- Kumar A, Yadav RP, Chatterjee S, Das M, Pal DK, et al., (2024) Integration of bioinformatics analysis to identify possible hub genes and important pathways associated with clear cell renal cell carcinoma. Urologia 91(2): 261-269.
- Oumarou Hama H, Aboudharam G, Barbieri R, Lepidi H, Drancourt M, et al., (2022) Immunohistochemical diagnosis of human infectious diseases: a review. Diagnostic Pathology 17(1): 17.
- Musa A, Ghoraie LS, Zhang SD, Glazko G, Yli-Harja O, et al., (2018) A review of connectivity map and computational approaches in pharmacogenomics. Brief Bioinform 19(3): 506-523.
- Li Y, L Li, (2019) Prognostic values and prospective pathway signaling of MicroRNA-182 in ovarian cancer: a study based on gene expression omnibus (GEO) and bioinformatics analysis. Journal of Ovarian Research 12(1): 106.
- Fabbro M, Lamy PJ, Touraine C, Floquet A, Ray-Coquard I, et al., (2023) HE4 and CA-125 kinetics to predict outcome in patients with recurrent epithelial ovarian carcinoma: the META4 clinical trial. Front Oncol 13: 1308630.
- Katsiki AD, Karatzas PE, De Lastic HX, Georgakilas AG,et al., (2024) DExplore: An Online Tool for Detecting Differentially Expressed Genes from mRNA Microarray Experiments. Biology (Basel) 13(5): 351.
- Alhumaydhi FA, (2022) Integrated computational approaches to screen gene expression data to determine key genes and therapeutic targets for type-2 diabetes mellitus. Saudi Journal of Biological Sciences 29(5): 3276-3286.
- Wolde T, Huang J, Huang P, Pandey V, Qin P, (2024) Depleted-MLH1 Expression Predicts Prognosis and Immunotherapeutic Efficacy in Uterine Corpus Endometrial Cancer: An In Silico Approach. BioMedInformatics 4(1): 326-346.
- Lu Y, Dai J, Liu Q, Cai P, Zheng J (2022) Screening of potential hub genes in pulmonary thromboembolism. Exp Ther Med 23(1): 18.
- Huang P, Wolde T, Bhardwaj V, Zhang X, Pandey V (2024) TFF3 and PVRL2 co-targeting identified by multi-omics approach as an effective cancer immunosuppression strategy. Life Sci 357: 123113.
- Liu W, Tu W, Li L, Liu Y, Wang S, et al., (2018) Revisiting Connectivity Map from a gene co-expression network analysis. Exp Ther Med 16(2): 493-500.
- Wei M, Bai X, Dong Q (2022) Identification of novel candidate genes and small molecule drugs in ovarian cancer by bioinformatics strategy. Transl Cancer Res 11(6): 1630-1643.
- Trott O, AJ Olson, (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2): 455-461.
- Chen Y, Ruben EA, Bieber MM, Rajadas J, Teng NH (2014) Abstract 5373: In silico investigation of FOXM1 binding and novel inhibitors in epithelial ovarian cancer (EOC). Cancer Research 74(19_Supplement): 5373-5373.
- Yang SYC, Lheureux S, Karakasis K, Burnier JV, Bruce JP, et al., (2018) Landscape of genomic alterations in high-grade serous ovarian cancer from exceptional long- and short-term survivors. Genome Med 10(1): 81.
- Gartel AL, (2017) FOXM1 in Cancer: Interactions and Vulnerabilities. Cancer Res 77(12): 3135-3139.
- Liu C, CJ Barger, AR Karpf (2021) FOXM1: A Multifunctional Oncoprotein and Emerging Therapeutic Target in Ovarian Cancer. Cancers (Basel) 13(12): 3065.
- Zona S, Bella L, Burton MJ, Nestal de Moraes G, Lam EWF (2014) FOXM1: An emerging master regulator of DNA damage response and genotoxic agent resistance. Biochimica et Biophysica Acta 1839 (11):1316-22.
- Huang YX, Zhao J, Song QH, Zheng LH, Fan C, et al., (2016) Virtual screening and experimental validation of novel histone deacetylase inhibitors. BMC Pharmacol Toxicol 17(1): 32.
- Bolton KL, Chen D, Corona de la Fuente R, Fu Z, Murali R, et al., (2022) Molecular Subclasses of Clear Cell Ovarian Carcinoma and Their Impact on Disease Behavior and Outcomes. Clin Cancer Res 28(22): 4947-4956.
- Abusharkh KAN, Onder FC, Çınar V, Hamurcu Z, Ozpolat B, et al., (2024) A drug repurposing study identifies novel FOXM1 inhibitors with in vitro activity against breast cancer cells. Med Oncol 41(8): 188.
- Zhang R, Siu MKY, Ngan HYS, Chan KKL (2022) Molecular Biomarkers for the Early Detection of Ovarian Cancer. International Journal of Molecular Sciences 23(19): 12041.
- Radu P, Kumar G, Cole A, Fameli A, Guthrie M, et al., (2024) Evolving assessment pathways for precision oncology medicines to improve patient access: a tumor-agnostic lens. Oncologist 29(6): 465-472.